Figures & data
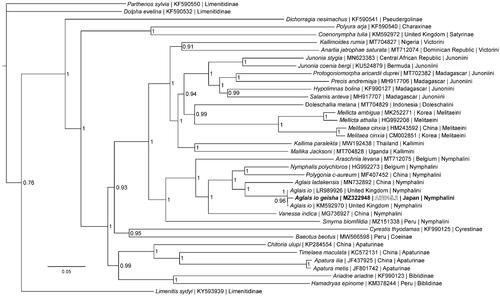
Figure 1. The Bayesian phylogeny (GTR + I + G model, average Potential Scale Reduction Factor (PSRF) = 1, average deviation of split frequencies = 0.000628) of the Aglais io geisha mitogenome, 37 additional mitogenomes from within family Nymphalidae, including outgroup species Limenitis sydyi, Parthenos sylvia, and Dophla evelina (Limenitinae) (Alexiuk et al. Citation2020b; Hamilton et al. Citation2020; Lalonde and Marcus Citation2020; Payment et al. Citation2020; Lalonde Citation2021), produced by 10 million MCMC generations in MrBayes, with sampling every 100 generations, and after discarding the first 250,000 generations as burn-in. The Bayesian posterior probability values determined by MrBayes are provided at each node.
Data availability statement
The genome sequence data that support the findings of this study are openly available in GenBank of NCBI at [https://www.ncbi.nlm.nih.gov] (https://www.ncbi.nlm.nih.gov/) under the accession nos. MZ322948 and MZ322949. The associated BioProject, SRA, and Bio-Sample numbers are PRJNA733565, SRX11064013, and SAMN19415664 respectively.
