Figures & data
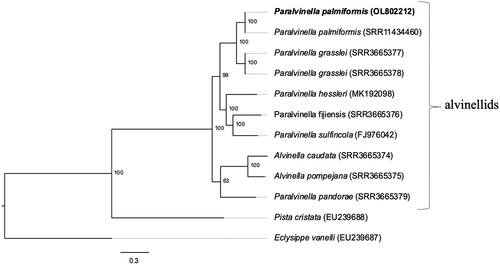
Figure 1. Maximum likelihood phylogeny (GTR nucleotide substitution model) using a 10,665 bp alignment of 12 mitochondrial protein coding genes. The outgroups are species of the polychaete families Terebellidae (P. cristata) and Amphraetidae (E. vanelli). The mitogenome sequenced in this study is indicated in bold. Numbers below nodes are the bootstrap values.
Data availability statement
This mitochondrial genome was deposited in GenBank of NCBI under the following accessions (BioProject: PRJNA786438, BioSample: SAMN23672631, Accession: OL802212). The P. palmiformis specimen and its DNA were deposited in the archives of the Deep-Sea Omics lab, Hong Kong University of Science and Technology under the accession P08H-3 (contact: Eric Lau, [email protected]). Additional data that support the findings of this study (e.g., gene sequence alignments) are openly available at https://github.com/maepz/Ppalmiformis_mitogenome.
