Figures & data
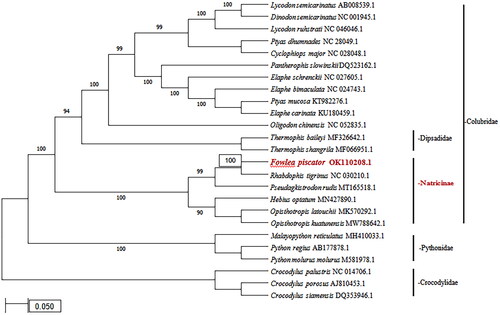
Figure 1. The Phylogenetic tree was constructed through MEGA 7.0 (Kumar et al. Citation2016) using the Maximum Likelihood method based on the Tamura-Nei model involving 25 complete mitochondrial genome sequences of Colubridae, Pythonidae and Crocodylidae species along with their accession numbers. All positions, containing gaps and missing data were eliminated. Bootstrap values are shown for each nodes based on 1000 replicates.
Data availability statement
The complete mitogenome sequence data that support the findings of this study are available in GenBank of NCBI (https://www.ncbi.nlm.nih.gov/nuccore/OK110208.1) under the accession no. OK110208.1. The associated BioProject, SRA and Bio-Sample numbers are PRJNA818295, SRR19137827 and SAMN27403394 respectively.
