Figures & data
Figure 1. The aerial stem of Zingiber striolatum (photo taken by Hong-Lei Li took at Xishuangbanna Botanical Garden, Jinghong, Yunnan).
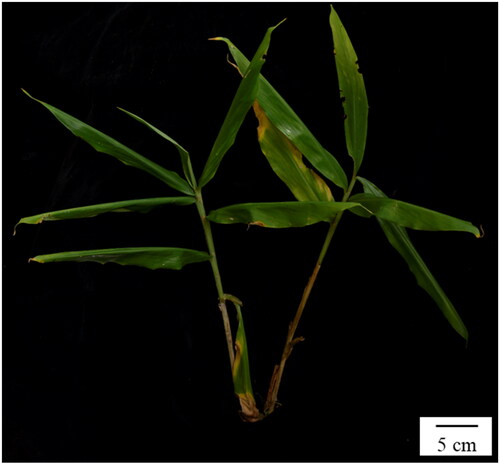

Figure 2. The structure of Zingiber striolatum chloroplast genome. The genes inside and outside of the circle are transcribed in the clockwise and counterclockwise directions, respectively. Genes belonging to different functional groups are shown in different colors. The darker gray area in the inner circle indicates the GC content and the lighter gray indicates the AT content of the chloroplast genome. The thick lines indicate the extent of the inverted repeats (IRA and IRB) that separate the chloroplast genome into the small single-copy (SSC) and large single-copy (LSC) regions.
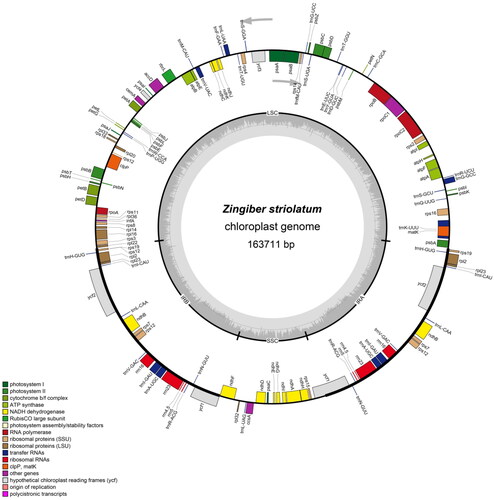
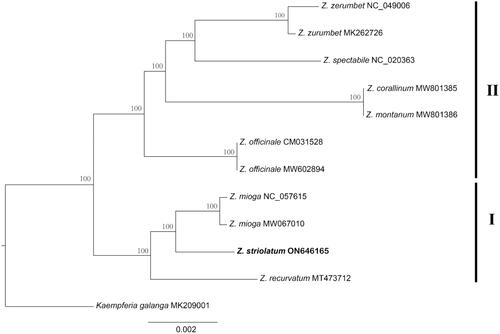
Figure 3. Phylogenetic tree of Zingiber inferred using maximum likelihood (ML) based on the complete chloroplast genome sequences. GenBank accession numbers: Z. striolatum (ON646165), Z. recurvatum (MT473712), Z. mioga (NC_057615), Z. officinale (MW602894), Z. montanum (MW801386), Z. corallinum (MW801385), Z. spectabile (NC_020363), Z. zerumbet (MK262726), and Kaempferia gaalanga (MK209001).
Data availability statement
The genome sequence data that support the findings of this study are openly available in GenBank of NCBI at (https://www.ncbi.nlm.nih.gov/nuccore/ON646165) under the accession no. ON646165. The associated BioProject, SRA, and Bio-Sample numbers are PRJNA842832, SRR19426750, and SAMN28688325 respectively.
