Figures & data
Figure 1. Sample collection of Camellia atrothea used for DNA sequencing in the present study. (A) Whole plant of the collected individual (imaged by Yanli Wang). (B) Mature leaves used for sequencing. (C) Statistics of the sequencing data.

Figure 2. The complete chloroplast genome map of C. atrothea. Arrangement of 133 genes represented in the map, including 88 protein coding genes, 37 tRNA genes, and 8 rRNA genes. The GC% along the chloroplast is represented by the inner circle.
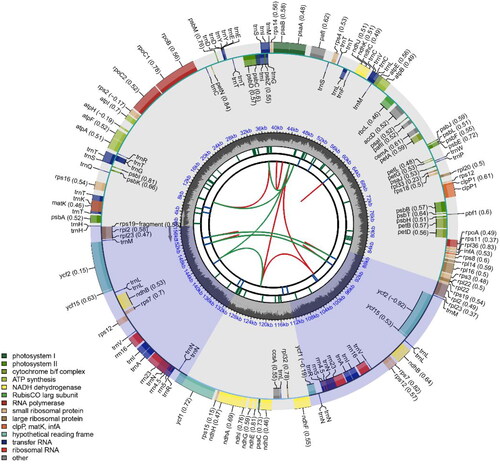
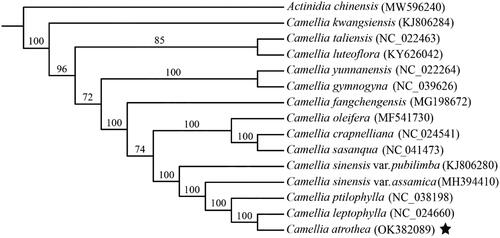
Figure 3. Maximum likelihood phylogenetic tree of C. atrothea and other 14 close plant species constructed using their complete chloroplast genome sequences. The bootstrap support value for each node is shown on the branch. The accession number of chloroplast genome of each plant species is shown in the brackets.
Supplemental Material
Download TIFF Image (14.1 MB)Supplemental Material
Download TIFF Image (4.1 MB)Supplemental Material
Download TIFF Image (15.1 MB)Supplemental Material
Download MS Word (240.7 KB)Supplemental Material
Download MS Excel (12.5 KB)Data availability statement
The assembled chloroplast genome sequence data that support the findings of this study are freely available in GenBank of NCBI (https://www.ncbi.nlm.nih.gov/) under the accession number of OK382089. The raw sequencing data of C. atrothea has also been deposited into the NCBI Bio-Project and SRA database with the accession number of PRJNA867579 and SRR20985353, respectively. The information of the plant material was deposited into NCBI Bio-Sample database with the accession number of SAMN30215288.
