Figures & data
Figure 3. Circular representation of Carallia brachiata chloroplast genome, showing the clockwise (genes inside the circle) and counterclockwise (outside) transcribed genes. Colors identify genes from the same functional category, following the figure legends. In the inner circle, the dark and light grey bars indicate the guanine + cytosine and adenine + thymine content, respectively. IRa and IRb: inverted repeat regions; LSC: large single copy region; SSC: small single copy.
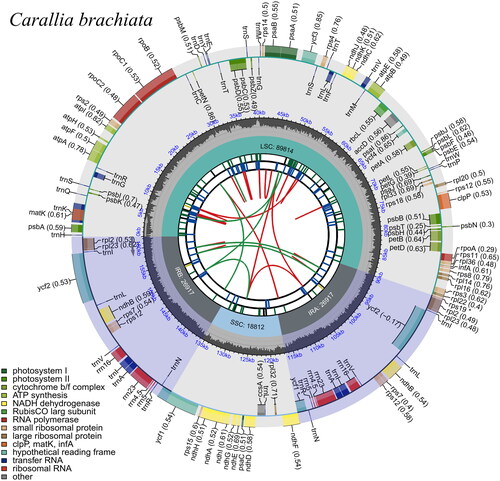
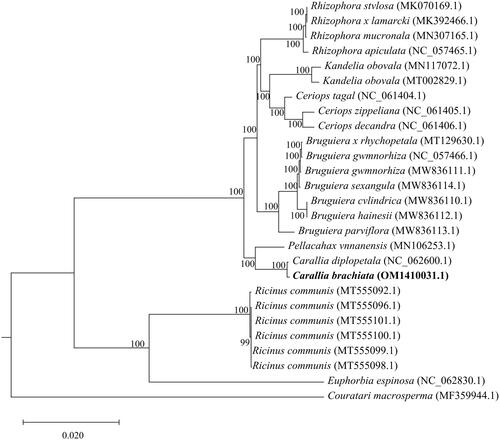
Figure 4. Maximum-likelihood phylogenetic tree for C. brachiata and 28 related species based on 62 homologous protein-coding genes. Bootstrap support values are indicated at each node (N = 1000). the scale bar indicates the phylogenetic distance in substitutions per site. The following sequences were used: Kandelia obovata (MN117072.1) (Du et al. Citation2019), Kandelia obovata (MT002829.1) (Xuli et al. 2020), Ceriops decandra (NC_061406.1) (Ruang-Areerate et al. Citation2022), Ceriops zippeliana (NC_061405.1) (Ruang-Areerate et al. Citation2022), Ceriops tagal (NC_061404.1) (Ruang-Areerate et al. Citation2022), Rhizophora stylosa (MK070169.1) (Li et al. Citation2019), Rhizophora × lamarcki (MK392466.1) (Zhang et al. Citation2019), Rhizophora mucronata (MN307165.1) (Wu Citation2019), Rhizophora apiculata (NC_057465.1) (Jiang Citation2020), Bruguiera gymnorhiza (NC_057466.1) (Jiang Citation2020), Bruguiera x rhynchopetala (MT129630.1) (Ying et al. Citation2020), Bruguiera gymnorhiza (MW836111.1) (Ruang-Areerate et al. Citation2021), Bruguiera sexangula (MW836114.1) (Ruang-Areerate et al. Citation2021), Bruguiera cylindrica (MW836110.1) (Ruang-Areerate et al. Citation2021), Bruguiera hainesii (MW836112.1)(Ruang-Areerate et al. Citation2021), Bruguiera parviflora (MW836113.1)(Ruang-Areerate et al. Citation2021), Carallia brachiata (OM141003.1) (this study), Carallia diplopetala (NC_062600.1) (Wang et al. Citation2021), Pellacalyx yunnanensis (MN106253.1) (Zhang et al. Citation2019), Ricinus communis (MT555096.1) (Muraguri et al. Citation2020), Ricinus communis (MT555101.1) (Muraguri et al. Citation2020), Ricinus communis (MT555100.1) (Muraguri et al. Citation2020), Ricinus communis (MT555099.1) (Muraguri et al. Citation2020), Ricinus communis (MT555098.1) (Muraguri et al. Citation2020), Ricinus communis (MT555092.1) (Muraguri et al. Citation2020), Euphorbia espinosa (NC_062830.1) (Wei Citation2021) and Couratari macrosperma (MF359944.1) (Vargas et al. Citation2017).
Supplemental Material
Download TIFF Image (61.4 KB)Supplemental Material
Download TIFF Image (23.2 KB)Supplemental Material
Download PDF (316.8 KB)Data availability statement
The genome sequence data that support the findings of this study are openly available in GenBank (https://www.ncbi.nlm.nih.gov/) under accession no. OM141003. The associated BioProject, SRA, and Bio-Sample numbers are PRJNA801906, SRR17823292, and SAMN25413005, respectively.