Figures & data
Figure 2. The chloroplast genome map of Cymbidium wenshanense. Genes drawn outside the outer circle were transcribed in a counterclockwise direction, and genes drawn inside the outer circle were transcribed clockwise. Genes that belong to different functional groups are color-coded. The colored legend at the bottom indicates genes with different functions. The dark grey inner circle indicates the GC content of the chloroplast genome and the presence of nodes in the LSC, SSC, and IR regions.
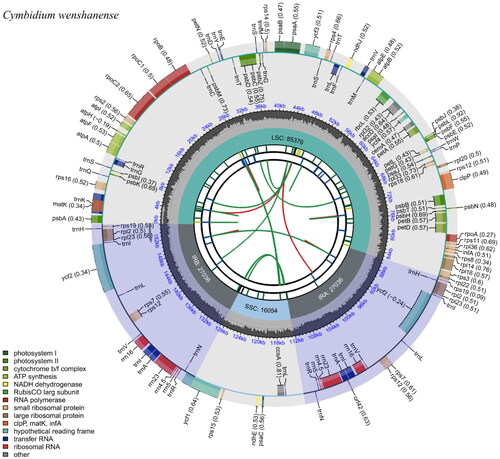
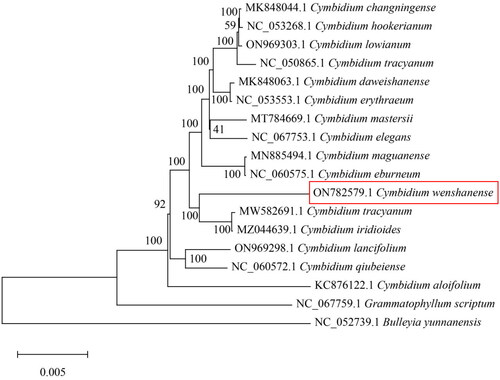
Figure 3. Phylogenetic tree based on 18 complete chloroplast genome sequences from Orchidaceae species. The following sequences were used: Cymbidium changningense (MK848044; Zheng et al. Citation2019), Cymbidium hookerianum (NC_053268), Cymbidium lowianum (ON969303), Cymbidium tracyanum (NC_050865), Cymbidium daweishanense (MK848063), Cymbidium erythraeum (NC_053553), Cymbidium mastersii (MT784669), Cymbidium elegans (NC_067753), Cymbidium maguanense (MN885494; Zheng et al. Citation2020), Cymbidium eburneum (NC_060575), Cymbidium tracyanum (MW582691; Zhe et al. Citation2022), Cymbidium iridioides (MZ044639), Cymbidium lancifolium (ON969298), Cymbidium qiubeiense (NC_060572.1), Cymbidium aloifolium (KC876122; Yang et al. Citation2013), Grammatophyllum scriptum (NC_067759), and Bulleyia yunnanensis (NC_052739).
Data availability statement
The data that support the findings of this study are openly available from the NCBI GenBank at https://www.ncbi.nlm.nih.gov, reference number ON782579.
The associated BioProject, SRA and Bio-Sample numbers are PRJNA764364, SRR15959874 and SAMN21500377, respectively.

