Figures & data
Figure 1. External features of P. liquidalis. This photo was taken by Hua Rong with the author’s approval for use.

Table 1. Species and GenBank accession number of mitogenomes used in this study.
Figure 2. Mitochondrial genome map of P. liquidalis. Represented with arrows, the transcription directions for the outer and inner genes are listed clockwise and anticlockwise, respectively.
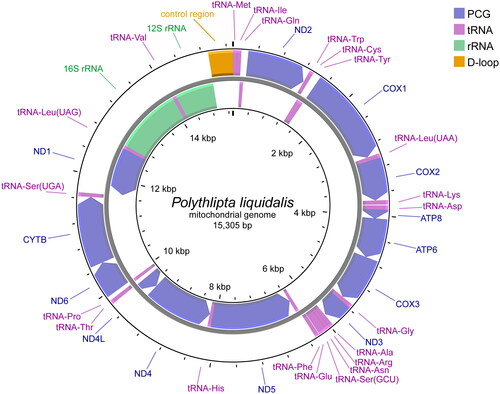
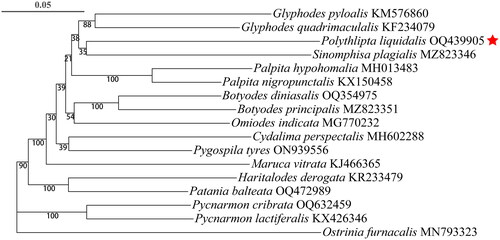
Figure 3. Phylogenetic trees using ML analyses based on 13 PCGs. The numbers above the branches are bootstrap support values (BS). alphanumeric terms indicate the GenBank accession numbers.

Supplemental Material
Download MS Word (566.5 KB)Data availability statement
The genome sequence data that support the findings of this study are openly available in GenBank of NCBI at (https://www.ncbi.nlm.nih.gov/) under the accession number OQ439905. The associated BioProject, SRA, and Bio-Sample numbers are PRJNA935109, SRR23461129, and SAMN33298022, respectively.
