Figures & data
Figure 1. The basidiocarp of Lanmaoa macrocarpa collected from Jiangxi Province, China. Photographed by Kuan Zhao.

Figure 2. The mitochondrial genome map of Lanmaoa macrocarpa. Genes outside and inside the outer circle are transcribed in counterclockwise and clockwise directions, respectively. Intron-contained genes were labeled with *. GC and at contents across the genome are shown with dark and light shading, respectively, inside the inner circle.
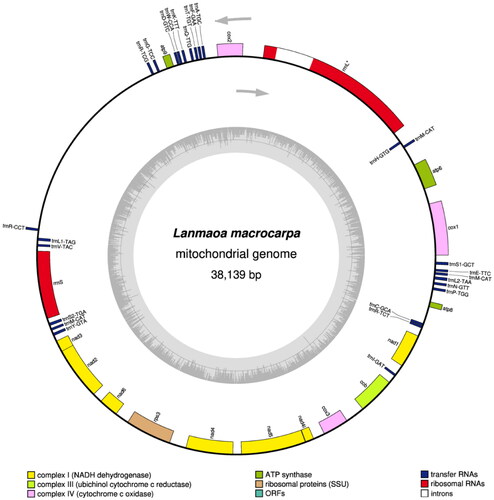
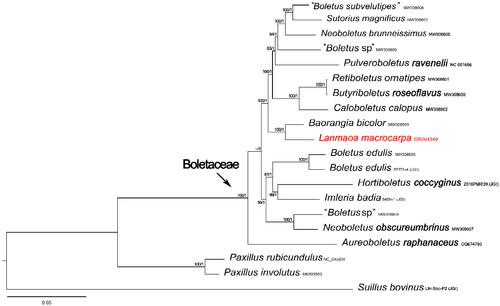
Figure 3. Phylogenetic tree of Lanmaoa macrocarpa and related taxa based on Bayesian inference (BI) and maximum likelihood (ML) analyses of 15 core protein coding genes. The GenBank accession numbers from NCBI or the information of voucher specimens from JGI followed the species names. The newly sequenced mitogenome is marked in red. Numbers near the nodes indicate bootstrap support values (>50%) and posterior probabilities (>0.95). the scale bar refers to 0.05 nucleotide substitutions per character.
Supplemental Material
Download PNG Image (222.5 KB)Data availability statement
The mitochondrial genome data is available with the accession number of OR004349 in the GenBank of NCBI (https://www.ncbi.nlm.nih.gov/). And the associated Bioproject, SRA, Bio-sample numbers are PRJNA957944, SRR24299300 and SAMN34273906, respectively.
