Figures & data
Figure 1. ‘Living cell of Haslea ostrearia observed in light microscopy’ from Gabed et al. (Citation2022).
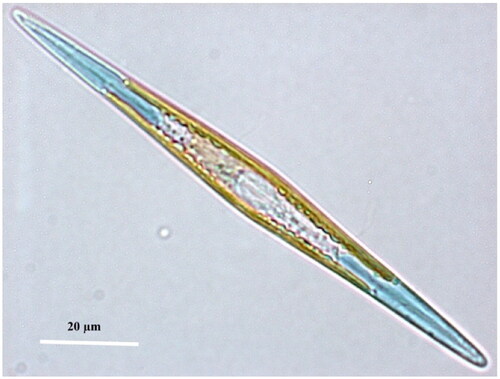

Figure 2. Genome map of the mitochondrial genome (A) and the chloroplastic genome (B) of H. ostrearia.
The red and blue colors in GC shiew (a) and GC plot (b) show if the value is below or above average. Protein-coding genes are shown in light blue, transfer RNA genes in light green and ribosomal RNA in light grey. The chloroplast genome is shown as a linear genome because it is not complete. The genome map was made using Artemis v18.2.0.
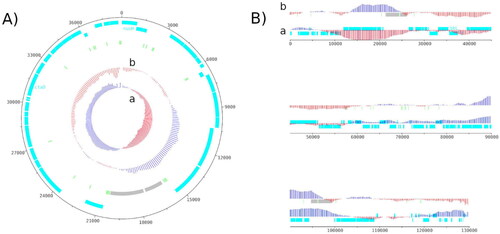
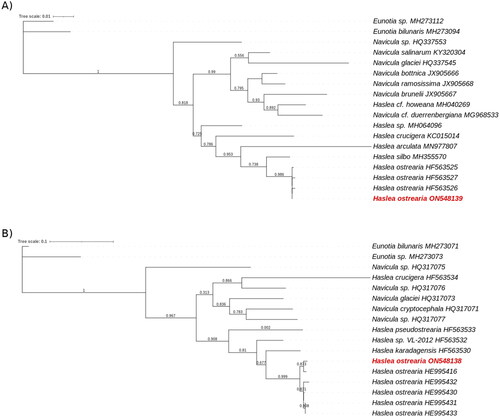
Figure 3. Maximum likelihood phylogenetic trees inferred from COX1 and rbcL genes from diatoms genus.
The phylogenetic trees was performed with, respectevily, Haslea ostrearia (in red) and 16 to 18 other diatom chloroplastic gene rbcL (A) and mictochondrial gene COX1 (B). Numbers near the nodes indicate bootstrap support values. The accession number associated with each gene is listed next to the species name. Eunotia species were used as external species. NGPhylogeny.fr pipeline was used to generate these phylogenetic trees.
Supplemental Material
Download PDF (293.9 KB)Data availability statement
The genome sequence data that support the findings of this study are openly available in GenBank of NCBI at https://www.ncbi.nlm.nih.gov/ under the accession no. ON548138 and ON548139. The associated BioProject, SRA and Bio-Sample numbers are PRJNA843895, SRR19450090/SRR19450089, and SAMN28772203 respectively.
