Figures & data
Figure 1. Species reference image of Poraniopsis inflata collected from Dokdo Island, Korea. Photographed by Taekjun Lee. This specimen in an ethyl alcohol solution (>95%) immediately after collected.

Figure 2. The complete mitochondrial genome of Poraniopsis inflata in this study. This map was generated by Geneious Prime ver. 2023.2.1.
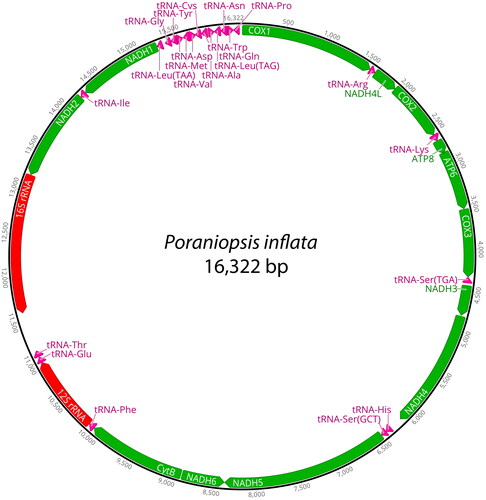
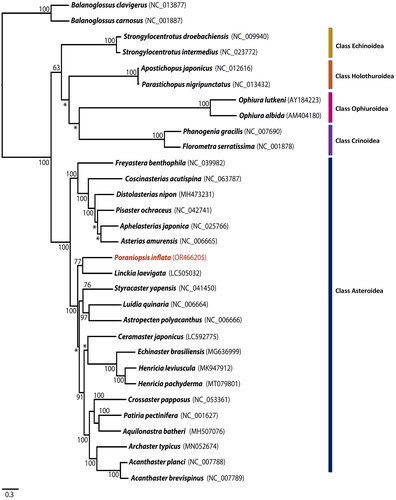
Figure 3. Phylogenetic analysis of Poraniopsis inflata and an additional 28 echinoderms was conducted using the maximum likelihood method based on the nucleotide sequences of 13 protein-coding genes. Two hemichordates, balanoglossus carnosus and B. clavigerus, were used as outgroups. The bootstrap support values are presented on each node, with values exceeding 50, while the asterisk marks indicate values below 50. Genbank accession numbers for published sequences are incorporated.
Supplemental Material
Download PNG Image (17.6 KB)Supplemental Material
Download PNG Image (24.5 KB)Data availability statement
BioSample, BioProject, and SRA accession numbers are https://www.ncbi.nlm.nih.gov/biosample/?term=SAMN37526385, https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1020671, and https://www.ncbi.nlm.nih.gov/sra/SRR26160490, respectively. The genome sequence data that support the findings of this study are openly available in GenBank of National Center for Biotechnology Information (NCBI) at https://www.ncbi.nlm.nih.gov, with an accession number OR466205.
