Figures & data
Figure 1. Specimen image of the sand bubbler crab, Scopimera longidactyla, collected from Daebu Island on the northwestern Coast of Korea. This photograph was taken by Dalyoung Kim in September 2022.

Table 1. List of accession number and publication used in the phylogenetic tree.
Figure 2. The circular mitogenome map of Scopimera longidactyla (GenBank Accession No. OR872329). Genes encoded on the reverse strand and forward strand are illustrated inside the circle and outside the circle, respectively.
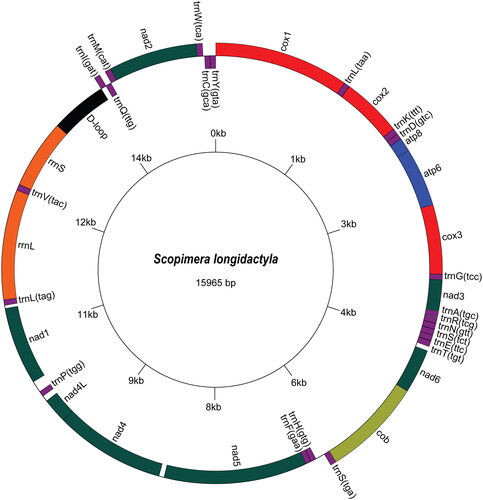
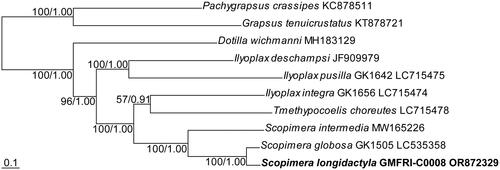
Figure 3. Maximum likelihood (ML) tree based on concatenated nucleotide sequences from all protein-coding genes of the complete mitogenomes of the family Dotillidae. Numeric values at nodes are ML bootstraps followed by the Bayesian posterior probabilities. The newly assembled mitogenome of the sand bubbler crab, Scopimera longidactyla was highlighted in bold. Two species, Pachygrapsus crassipes and Grapsus tenuicrustatus, belonging to the family Grapsidae, were selected as the outgroup. Scale bar represents nucleotide substitutions per site.

Supplemental Material
Download PDF (206.2 KB)Data availability
The data that support the findings of this study are openly available in GenBank of NCBI under the accession number OR872329. Raw reads have been deposited under NCBI BioProject (PRJNA1041591), BioSample (SAMN38288372), and SRA (SRR26857270).
