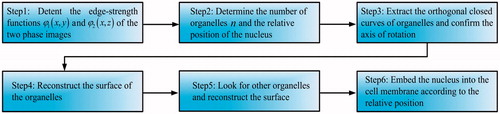
Figures & data
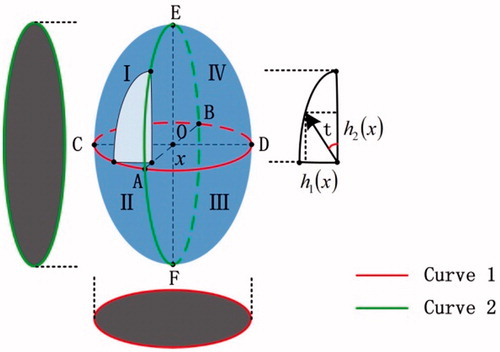
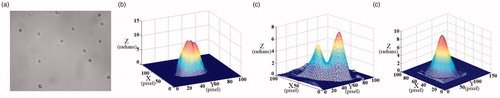
Figure 1. (a) Binucleate cell micrograph; (b) the approximate model; (c,d) the 2D and 3D phase distribution along y axis; (e,f) the 2D and 3D phase distribution along z axis.
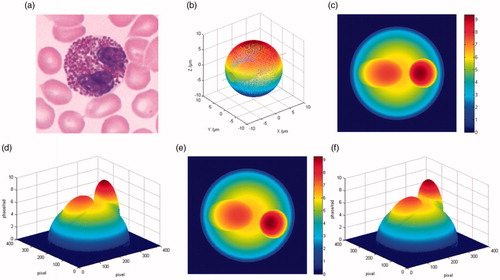
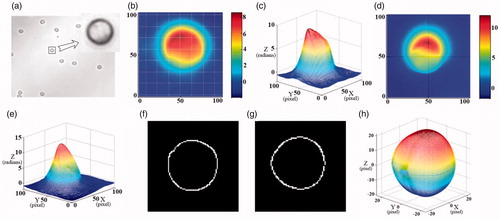
Figure 4. Binuclear cell contour images. (a) The edge-strength of the phase image in x-z plane, (b–d) the contours of the cell membrane, ellipsoidal nucleus and spherical nucleus, (e–h) the corresponding results in x-y plane.
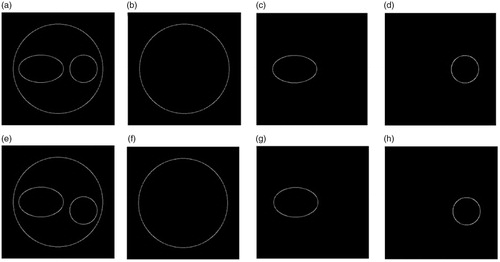

Figure 5. Dinuclear cell reconstruction results by geometric rotation algorithm. (a–c) The 3D surfaces of the cell membrane, ellipsoidal nucleus and spherical nucleus; (d) 3D complete surface of the cell.
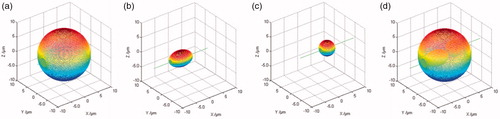
Table 1. Reconstruction errors of each part of the cell.
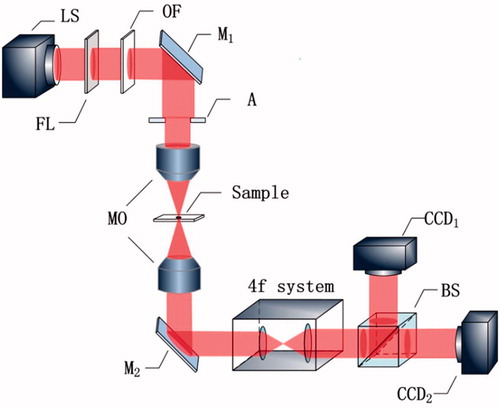
Figure 6. Principle of experimental setup. LS: light source; FL: frosted lens; OF: optical filter; A: aperture; M: mirror; MO: microscopy objective; BS: beam splitter.