Figures & data
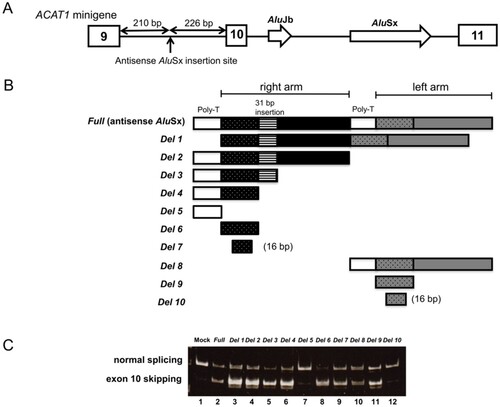
Figure 1. Splicing assays using the ACAT1 minigene with AluSx deletions. (A) Schematic of the ACAT1 minigene including exon 9 to exon 11 of ACAT1. White boxes and white arrows indicate exons and the original Alu elements, respectively. Full AluSx or AluSx deletions were inserted in an antisense orientation into ACAT1 intron 9 at 226 bp upstream of exon 10. (B) The structure of full-length AluSx and the 10 antisense AluSx deletion constructs. (C) Minigene splicing experiments with the deletion constructs. Mock experiments were performed with an ACAT1 minigene construct without antisense AluSx. Normal splicing (exon 10 inclusion) and aberrant splicing (exon 10 skipping) generated 309 and 244 bp fragments, respectively.
Supplemental Material
Download Zip (277.3 KB)Data availability statement
The data that support the findings of this study are available at the figshare repository (https://figshare.com/) at https://doi.org/10.6084/m9.figshare.16531881.v1. Nucleotide sequences of antisense AluSx in this study are available at https://doi.org/10.6084/m9.figshare.16531659.

