Figures & data
Figure 1. S-Nitrosylation of p47phox. (A) Immunoblot analysis of recombinant p47phox. After incubating p47phox with various concentrations of SNAP or PN for 30 minutes at room temperature, the samples were subjected to SDS–PAGE and immunoblotting with the anti-nitrocysteine IgG antibody. (B) Identification of the S-nitrosylation target cysteine in p47phox by using site-directed mutagenesis. The S-nitrosylation of wild type and p47phox C→A mutants was detected using the DAN reagent. The fluorescence signal of the untreated control protein was subtracted as background. Values correspond to three independent assays with the standard deviation indicated. *P < 0.01 versus wild type, C98A, C111A, and C396 p47phox mutant proteins. (C) After incubation with 1 mM SNAP or 1 mM PN for 30 minutes at room temperature, the wild type and the p47phox C196A mutant were subjected to SDS–PAGE, followed by immunoblotting with the anti-nitrocysteine IgG antibody.
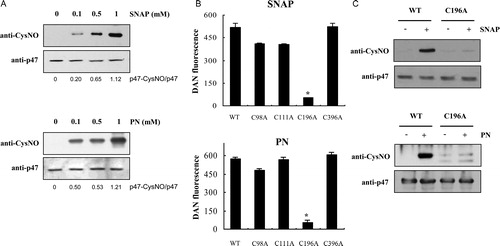
Figure 2. Effect of S-nitrosylation on in vitro phosphorylation of recombinant p47phox by CK2. (A) Recombinant p47phox (1 μg) was incubated with various concentrations of SNAP or PN (30 minutes at room temperature), followed by incubation with [gamma-32P]ATP and CK2. The labeled proteins were separated by SDS–PAGE. Proteins were visualized using the anti-p47phox IgG antibody, and 32P incorporation (p-p47phox) was monitored by autoradiography. (B) Effect of 1 mM DTT and degraded SNAP or PN on the phosphorylation of S-nitrosylated p47phox by CK2. (C) SNAP- or PN-treated wild type and p47phox C196A mutant samples were phosphorylated by CK2 with [gamma-32P]ATP. The labeled proteins were separated by SDS–PAGE, and 32P incorporation was monitored by autoradiography. (D) Recombinant p47phox (1 μg) was incubated with various concentrations of PN (30 minutes at room temperature) and then with [gamma-32P]ATP and PKC. The labeled proteins were separated by SDS–PAGE. Proteins were visualized with anti-p47phox IgG, and 32P incorporation was monitored by autoradiography.
![Figure 2. Effect of S-nitrosylation on in vitro phosphorylation of recombinant p47phox by CK2. (A) Recombinant p47phox (1 μg) was incubated with various concentrations of SNAP or PN (30 minutes at room temperature), followed by incubation with [gamma-32P]ATP and CK2. The labeled proteins were separated by SDS–PAGE. Proteins were visualized using the anti-p47phox IgG antibody, and 32P incorporation (p-p47phox) was monitored by autoradiography. (B) Effect of 1 mM DTT and degraded SNAP or PN on the phosphorylation of S-nitrosylated p47phox by CK2. (C) SNAP- or PN-treated wild type and p47phox C196A mutant samples were phosphorylated by CK2 with [gamma-32P]ATP. The labeled proteins were separated by SDS–PAGE, and 32P incorporation was monitored by autoradiography. (D) Recombinant p47phox (1 μg) was incubated with various concentrations of PN (30 minutes at room temperature) and then with [gamma-32P]ATP and PKC. The labeled proteins were separated by SDS–PAGE. Proteins were visualized with anti-p47phox IgG, and 32P incorporation was monitored by autoradiography.](/cms/asset/f455bb9a-d212-49ef-a769-44000c66fa62/yrer_a_11646844_f0002_b.jpg)
Figure 3. Conformational change in p47phox protein by S-nitrosylation. (A) Recombinant wild type and SH3 domains of p47phox (1 μg) were incubated with [gamma-32P]ATP and CK2. The labeled proteins were separated by SDS–PAGE. Proteins were visualized using the anti-p47phox IgG antibody, and 32P incorporation was monitored by autoradiography. (B) The SH3 domain of p47phox (1 μg) was incubated with PN (30 minutes at room temperature) and then with [gamma-32P]ATP and CK2. The labeled proteins were separated by SDS–PAGE. Proteins were visualized using the anti-p47phox IgG antibody, and 32P incorporation (p-SH3) was monitored by autoradiography. (C) Changes in p47phox structure following S-nitrosylation. Steady-state emission spectra of intrinsic fluorescence of native p47phox and SH3 domains treated with SNAP or PN were analyzed with a spectrophotometer at an excitation wavelength of 278 nm. The maximum protein fluorescence intensity was determined at 338 nm. Data are representative of three experiments. *P < 0.01 versus untreated p47phox. (D) Visualization of binding of CK2-beta subunit to S-nitrosylated p47phox using dot blot analysis.
![Figure 3. Conformational change in p47phox protein by S-nitrosylation. (A) Recombinant wild type and SH3 domains of p47phox (1 μg) were incubated with [gamma-32P]ATP and CK2. The labeled proteins were separated by SDS–PAGE. Proteins were visualized using the anti-p47phox IgG antibody, and 32P incorporation was monitored by autoradiography. (B) The SH3 domain of p47phox (1 μg) was incubated with PN (30 minutes at room temperature) and then with [gamma-32P]ATP and CK2. The labeled proteins were separated by SDS–PAGE. Proteins were visualized using the anti-p47phox IgG antibody, and 32P incorporation (p-SH3) was monitored by autoradiography. (C) Changes in p47phox structure following S-nitrosylation. Steady-state emission spectra of intrinsic fluorescence of native p47phox and SH3 domains treated with SNAP or PN were analyzed with a spectrophotometer at an excitation wavelength of 278 nm. The maximum protein fluorescence intensity was determined at 338 nm. Data are representative of three experiments. *P < 0.01 versus untreated p47phox. (D) Visualization of binding of CK2-beta subunit to S-nitrosylated p47phox using dot blot analysis.](/cms/asset/4087fd2e-7165-42f4-8175-5a5e2420df3d/yrer_a_11646844_f0003_b.jpg)