Figures & data
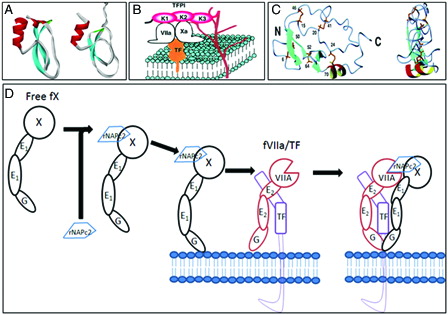
Figure 1. (A) Ribbon diagrams of the second and third Kuniz domain of TFPI. (B) Mechanism of action of TFPI: inhibitory mechanisms of the initial reaction of blood coagulation by TFPI on cell surface. TFPI binds to FVIIa and Xa via K1 and K2 domains and to proteoglycans via K3 and C-terminal domains (K denotes Kunitz domain) (diagram kindly provided by Dr Jun Mizuguchi). (C) Ribbon diagram of the minimized mean structures of NAPc2 (kindly provided by Dr Peter E Wright). The molecules on the right have been rotated about the vertical axis by 90°. Beta-Strands are shown in cyan and helices in red and yellow. The cysteine side chains are shown in gold. (D) Mechanism of action of rNAPc2. In step 1 of the scheme, rNAPc2 binds with high affinity to zymogen FX in solution (binding to FXa as an inhibitory scaffold is not shown). The resulting stable bimolecular complex between rNAPc2–FX serves as an inhibitory scaffold that facilitates docking to the membrane-bound TF–FVIIa. In this way, the inhibitory scaffold uses both a similar docking mechanism as the natural substrate (FX) and subsequent presentation to the active site of TF–FVIIa. In the last step of the mechanistic scheme, the canonical reactive loop of rNAPc2 docks into the active site of FVIIa/TF, allowing for the formation of tightly bound quaternary inhibitory complex.