Figures & data
List of all observed testate amoebae taxa. Codes used in are added
APPENDIX 1 (Cont.)
FIGURE 1. Sketch map of Antarctica, the Crozet Archipelago, and Île de la Possession showing the locations of the different sampling sites (only first and last sample numbers are shown)
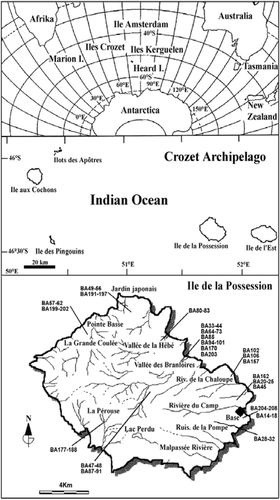
FIGURE 2. Cumulative species diversity for samples, shown as the mean, ± 2 standard errors. Standard errors are based on all samples
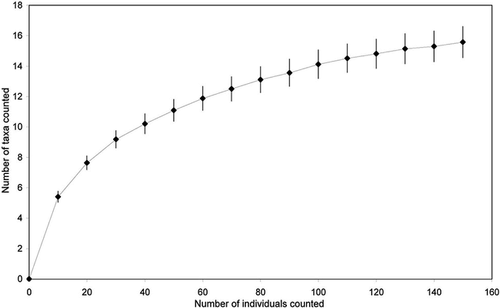
FIGURE 3. Relative abundance of the main Testacea genera observed in this study (black bars) and the number of taxa occurring in the different genera (white bars)
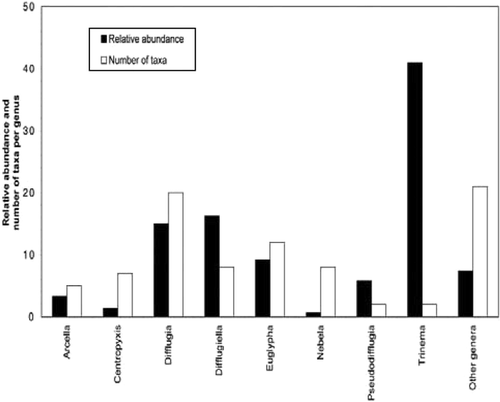
FIGURE 4. Cluster dendrogram showing all samples used for analysis (66). Letters indicated on the right refer to the different assemblages: (A) Difflugiella crenulata assemblage; (B) Trinema assemblage. Lower axis shows the linkage distance
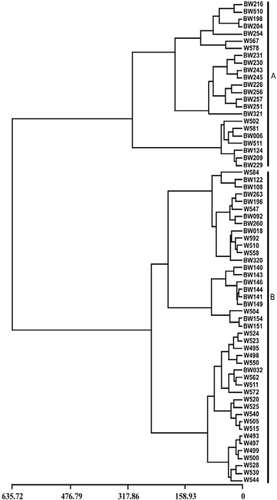
FIGURE 5. Redundancy Analysis (RDA) ordination diagram showing the relationship between sites and environmental variables. Only RDA axes 1 and 2 are shown. Arrows indicate environmental variables as detected by forward selection. Continuous variables are represented by arrows, categorical variables are indicated with a star. Samples labelled with and open circle belong to the Trinema assemblage; samples with an open triangle belong to the Difflugiella crenulata assemblage
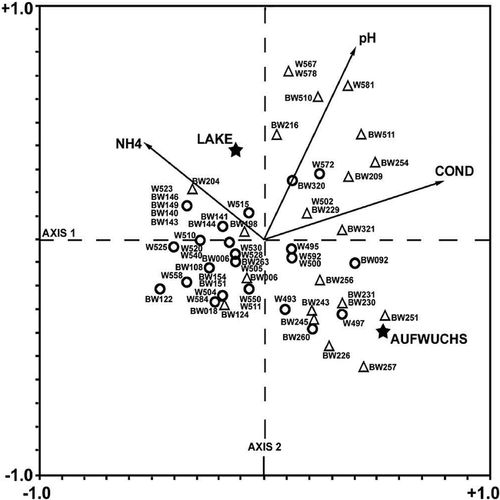
FIGURE 6. Plot of observed versus inferred pH values. Linear regression equation is y = 0.9999x + 0.0006, where x and y are the observed and the calculated pH values, respectively. The dark diamonds represent the different samples
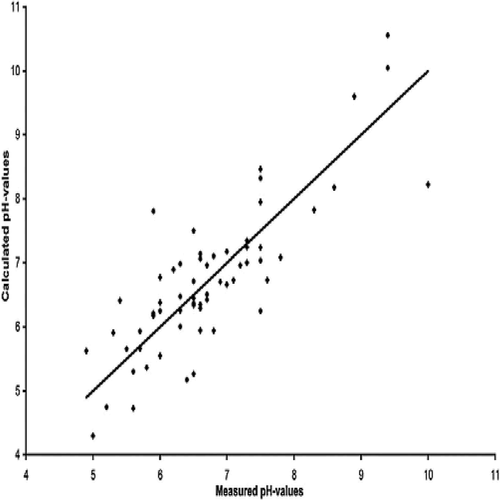
FIGURE 7. Plot of the observed pH values versus the residulas. Linear regression equation is y = 0.0002x − 0.0001 where x represents the observed pH values and y the residuals. The dark diamonds represent the different samples
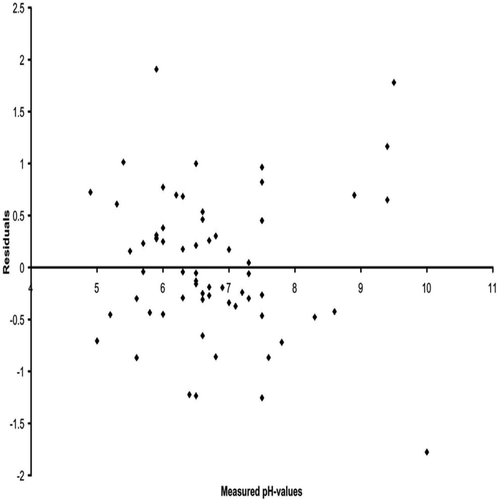
FIGURE 8. Weighted Average (WA) optima and tolerances for pH illustrated in ascending order for the principal testate amoebae species used to develop the model. Codes are explained in Appendix 1
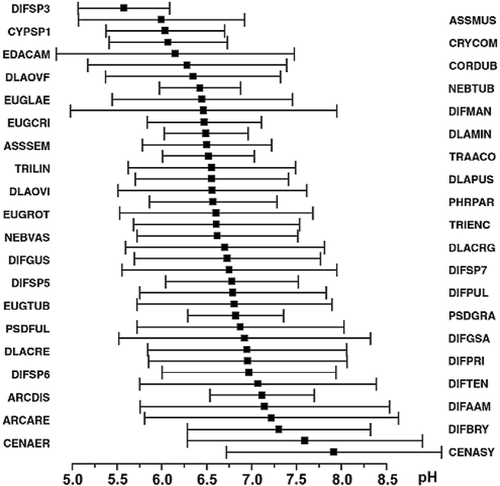
TABLE 1 Some characteristics of the different assemblages (mean numbers ± standard errors)