Figures & data
Table 1 Specific activity of family I (Rsp. rubrum) and family II (Rba. sphaeroides, Rba. capsulatus, Rdv. sulfidophilum) PPases, with and without divalent cation added at the hydrolysis reaction medium: 50 mM Tris–HCl (pH 8.6), 2 mM Na4PPi, and 3 mM of divalent cation
Figure 1 EDTA effect on the hydrolysis of pyrophosphatase.
Abbreviations: EDTA, ethylenediaminetetraacetic acid; Rsp. rubrum, Rhodospirillum rubrum ;Rba. capsulatus, Rhodobacter capsulatus ;Rba. sphaeroides, Rhodobacter sphaeroides ;Rdv. sulfidophilum, Rhodovulum sulfidophilum.
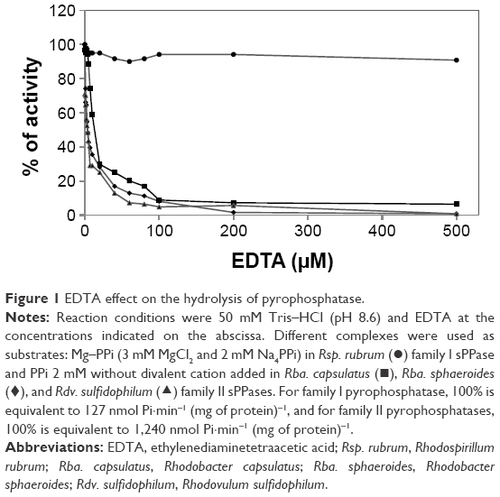
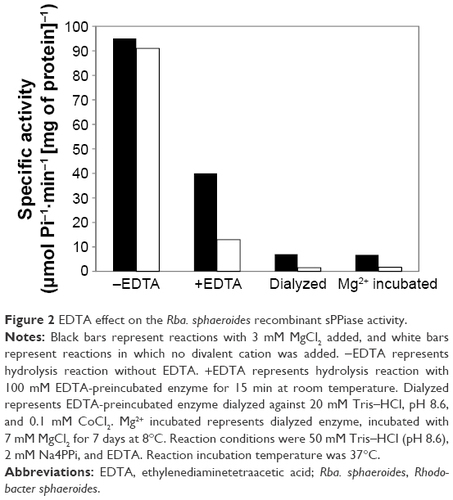
Figure 2 EDTA effect on the Rba. sphaeroides recombinant sPPiase activity.
Abbreviations: EDTA, ethylenediaminetetraacetic acid; Rba. sphaeroides, Rhodobacter sphaeroides.