Figures & data
Table 1 Antimalarial Drug Mechanism and Resistance Marker
Figure 1 Major antimalarial drugs and their tolerance. (A) Hemozoin formation in food vacuoles: (i) 4- aminoquinoles including chloroquine (CQ), amodiaquine (AQ), piperaquine (PPQ), (ii) Amino alcohols including mefloquine (MFQ), lumefantrine (LMF), quinine (QN) and (iii) Artemisinin (ART) bind to reactive heme and interfere its detoxification by making a complex with heme, thus inhibiting the formation of hemozoin. QN and MFQ are sensitive to PfCRT and PfMDR1. Resistance to these drugs occurs by drug mutation and copy number variation in pfcrt and pfmdr1 gene (shown by red diamond). (B) Folate synthetic in the cytosol: Folate synthetic pathway enzymes dihydropteroate synthase is sensitive to the drug sulfadoxine (SDX) and dihydrofolate reductase is sensitive to pyrimethamine (PYR) and proguanil (PRG). Resistance to SDX and PYR occurs due to mutation (shown by red diamond) in dhps and dhfr genes, respectively. (C) Electron transport chain in mitochondria: Atovaquone (ATQ) and ART inhibit cytochrome b complex (CyTb) in the electron transport chain by generating reactive oxygen species. ART in mitochondria causes protein and lipid alkylation, leading to oxidative stress and cellular damage. Resistance to these drugs is caused by a mutation (shown by red diamond) in the cytb gene. (D) Protein synthesis in apicoplast: antibiotics doxycycline (DOX), clindamycin (CLD), and azithromycin (AZT) inhibit protein synthesis by targeting the ribosomal RNA of the apicoplast. CLD resistance is mediated by a point mutation (shown by red diamond) in the apicoplast-encoded 23S rRNA. (E) Endoplasmic reticulum: ART is sensitive to PfATP6 in the endoplasmic reticulum. Variants in Kelch 13 (K13) are the primary mediator ring-stage parasite resistance to ART (shown by red diamond).
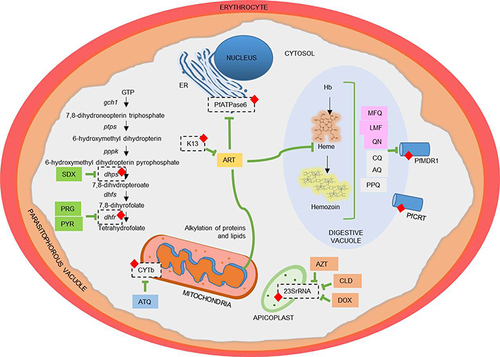
Figure 2 Timeline of chloroquine resistance in Indonesia. This figure summarizes the development of chloroquine resistance in Indonesia from 1974 until 2015 based on their locations and identification methods.
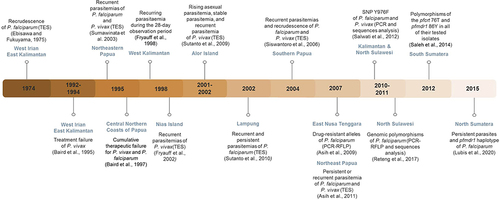
Figure 3 Timeline of sulfadoxine–pyrimethamine resistance in Indonesia. The development of sulfadoxine–pyrimethamine resistance in Indonesia was documented from 1979 until 2010. In each instance, the locations and identification methods of resistance are summarized.
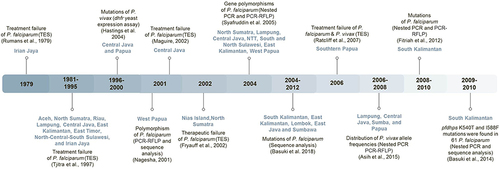
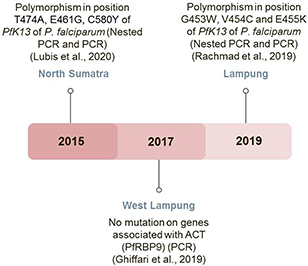
Figure 4 Identification of alleles mutant in PfK13 as an early warning sign for artemisinin resistance in Indonesia. The use of artemisinin combination therapy as a first-line antimalarial drug in Indonesia was summarized from 2015–2019. The polymorphisms of PfK13 serve as a warning sign of resistance toward this drug.