Figures & data
Figure 1 The levels of MOTS-c and experimental behavioral results of mice in different groups (n=7).
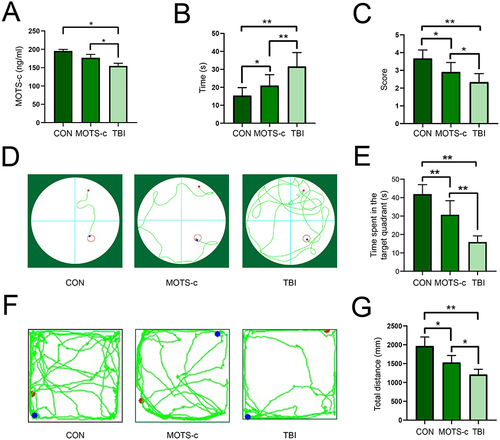
Figure 2 Transcriptomics analysis of MOTS-c VS TBI groups (n=3).
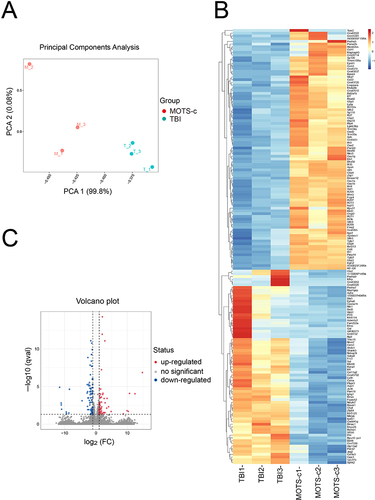
Figure 3 GO terms and KEGG pathway analyses of the DEGs (n=3).
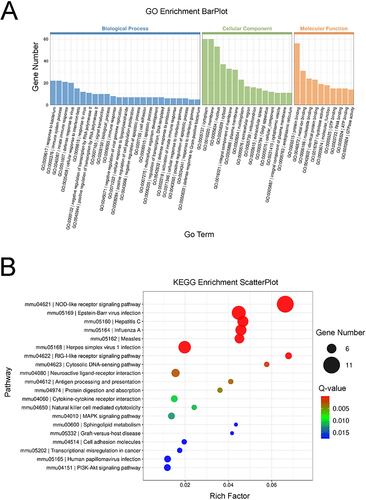
Figure 4 Metabolomics analysis of MOTS-c VS TBI groups (n=7).
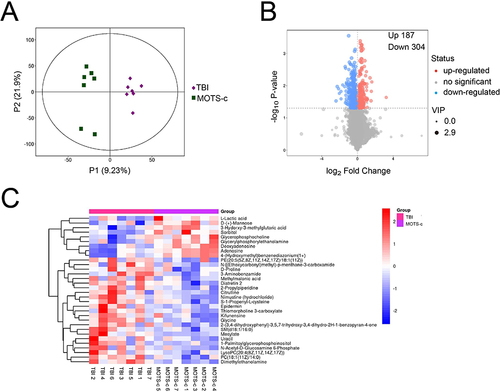
Figure 5 Pathway analysis of the DEMs (n=7).
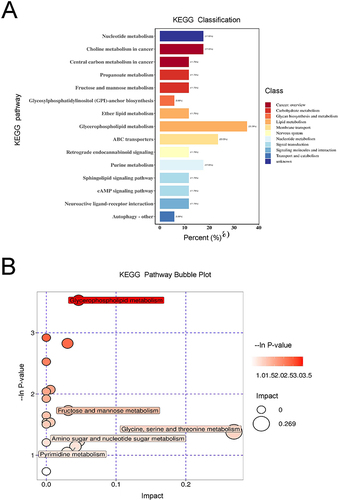
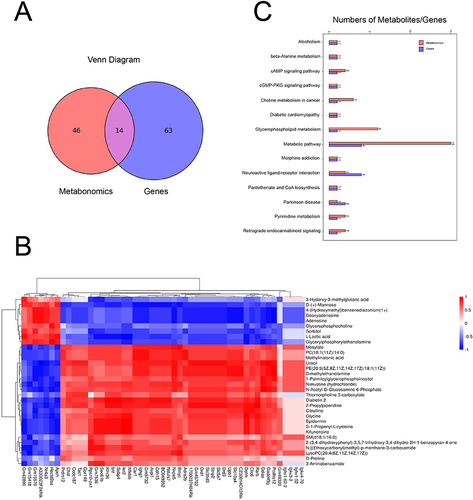
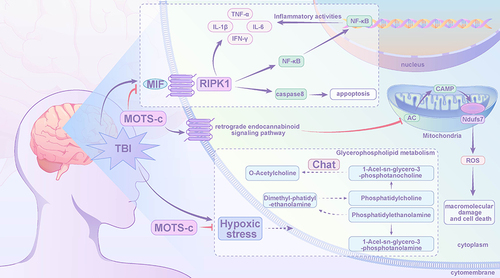
Figure 6 Integrated analysis of metabolomics and transcriptomics data.