Figures & data
Figure 1 Overview of the HIF system.
Abbreviations: PHD, Prolyl hydroxylases; HIF, hypoxia-inducible transcription factor; ODD, oxygen-dependent degradation domain; bHLH, basic helix-loop-helix motif; PAS, Per-ARNT-Sim domain; CTAD, C-terminal transactivation domain; PAC, PAS-associated C-terminal domain; FIH, factor inhibiting HIF.
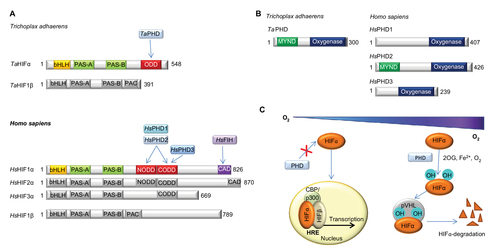
Figure 2 Evidence for conservation of biochemical properties between TaPHD and HsPHD2.
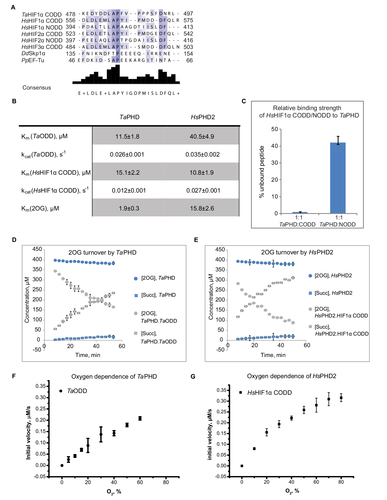
Table 1 Data collection and refinement statistics
Figure 3 Views from crystal structures of the Trichoplax adhaerens prolyl hydroxylases (PHD) without substrate bound (TaPHD) and in complex with a fragment of its substrate (TaPHD.TaODD).
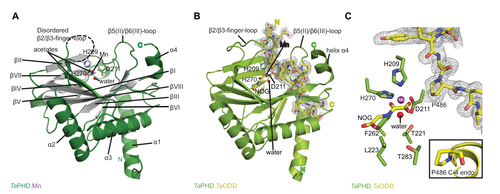
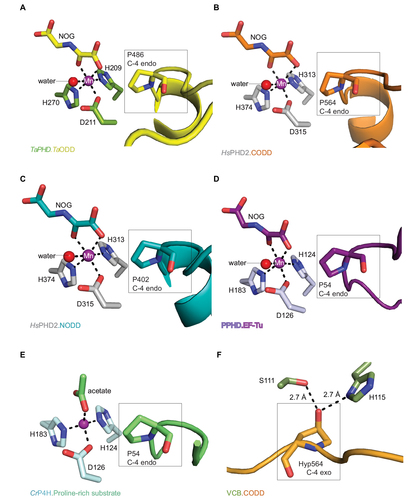
Figure 4 Comparison of ODD binding modes by the T. adhaerens and human HIFα PHD for the TaPHD.TaODD, HsPHD2.CODD and HsPHD2.NODD substrate structures.
Abbreviations: T. adhaerens, Trichoplax adhaerens; ODD, oxygen dependent degradation domain.
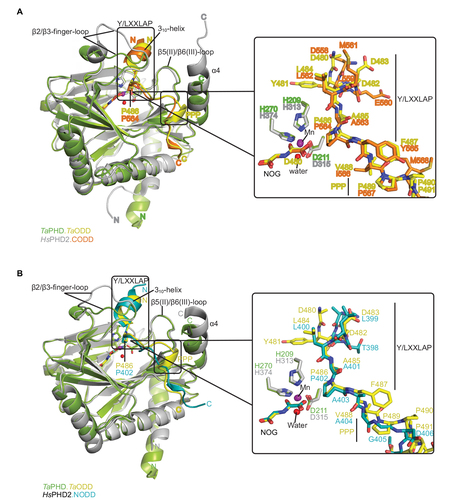
Figure 5 Comparison of substrate binding modes by TaPHD, CrP4H and Pseudomonas putida PPHD.
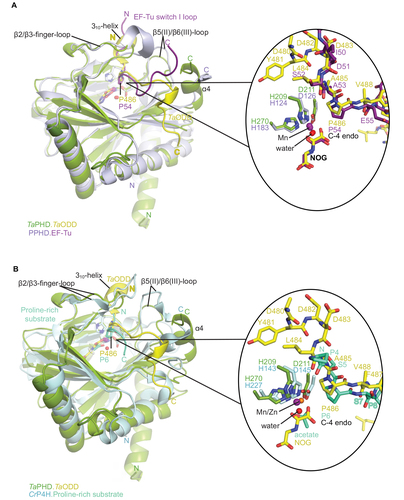
Figure 6 Active site metal region details and comparison of the conformations of the target prolyl-residues in the enzyme-substrate complex structures of TaPHD, HsPHD2, PPHD, and CrP4H.