Figures & data
Table 1 Characteristics of 52 Patients with Elizabethkingia Colonization or Infection
Figure 1 The distribution of 52 Elizabethkingia isolates according to the year and site of isolation.
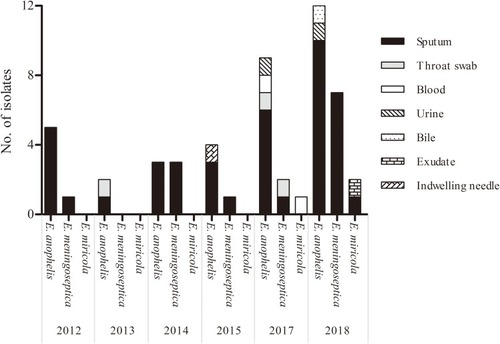
Table 2 Antimicrobial Susceptibilities of 52 Elizabethkingia Isolates Determined by the Broth Microdilution Method
Table 3 Distribution of β-Lactamase Genes in 52 Elizabethkingia
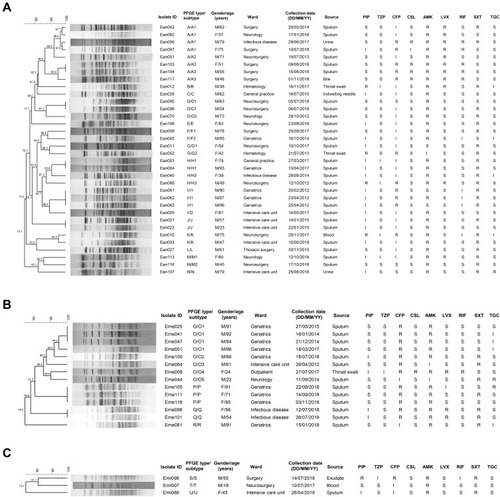
Figure 2 Dendrogram of PFGE patterns of 52 Elizabethkingia isolates using the BioNumerics software. (A) Thirty-four E. anophelis isolates; (B) Fourteen E. meningoseptica isolates; (C) Three E. miricola isolates.
Abbreviations: M, male; F, female; PIP, piperacillin; TZP, piperacillin-tazobactam; CFP, cefoperazone; CSL, cefoperazone-sulbactam; AMK, amikacin; LVX, levofloxacin; RIF, rifampin; SXT, trimethoprim-sulfamethoxazole; TGC, tigecycline; S, susceptible; I, intermediate; R, resistant.

