Figures & data
Table 1 Primers and Cyclic Conditions Used in PCR Targeting Three Classes of Carbapenemases and BoxA Region Among CRKP Clinical Isolates
Figure 1 Phenotypic tests for detection and characterization of carbapenemase production. (A); Modified Hodge test (K02 to K17 denotes to isolate code), (B); Carba NP test (row A; denotes to solution A, row B; denotes to solution B, row C; denotes to solution A containing tested isolates, rows D & E; denote to solution B containing tested isolates, red color indicates negative result, yellow color indicates positive result) (C); Combined disk synergy test (disk A; meropenem, disk B; meropenem/clavulanic acid, disk C; meropenem/sulbactam, disk D; meropenem/tazobactam, disk E; meropenem/phenylboronic acid, disk F; meropenem/EDTA, disk G; meropenem/mercaptopropionic acid, disk H; meropenem/sodium chloride).
Abbreviations: NP, Nordmann/Poirel; EDTA, ethylenediaminetetraacetic acid.
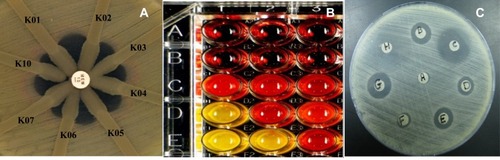
Table 2 Antimicrobial Susceptibility Pattern of CRKP Clinical Isolates
Table 3 Phenotypic and Genotypic Screening Profiles of CRKP Clinical Isolates
Table 4 Carbapenemase Inhibitor Profile of CRKP Clinical Isolates
Table 5 Detailed Resistance and Genetic Profiles of CRKP Clinical Isolates
Figure 2 Multiple sequence alignment of sequenced blaNDMwith the reference sequence blaNDM-1 KP036459.1; the arrow (A) is pointing to thymine nucleotide in reference sequence blaNDM-1 KP036459.1; the arrow (B) is pointing to non-mutated thymine nucleotide at position 558 in the query sequence of investigated blaNDM-1; the arrow (C) is pointing to mutated guanine nucleotide at position 558 in the query sequence of investigated blaNDM-25.
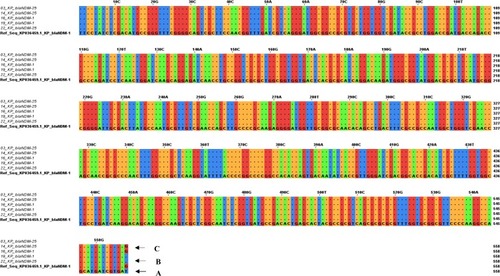
Figure 3 Representative DNA fingerprint pattern generated by BOX-PCR for carbapenem-resistant K. pneumoniae clinical isolates. L, 100 bp DNA ladder; lanes 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, and 18 are the code No. of CRKP clinical isolates.
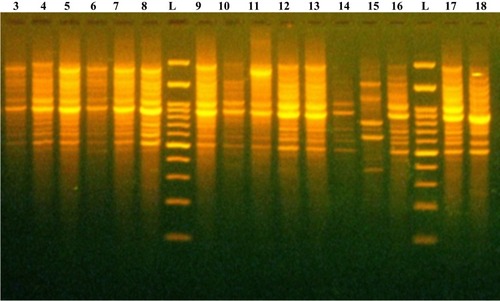
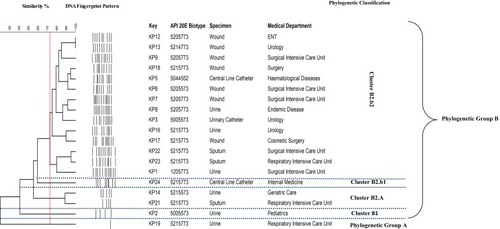
Figure 4 Unweighted pair-group method with arithmetic average (UPGMA) dendrogram based on Dice similarity for banding profile generated by BOX-PCR.