Figures & data
Figure 1 Sanger sequencing for validation of the variations detected by the next-generation sequencing platforms. (A) ATP6V0A4 c.1418C>T variant in family members of Individual 1. (B) POU1F1 c.416G>A (p. Arg139Gln) variant in family of Individual 1.
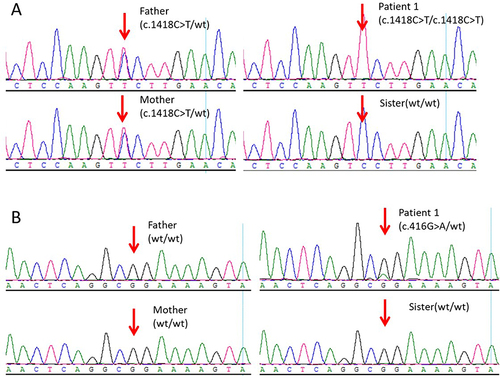
Figure 2 Sanger sequencing for validation of the variations detected by the next-generation sequencing platforms. POU1F1 c.212C>T variant in family members of Individual 2.
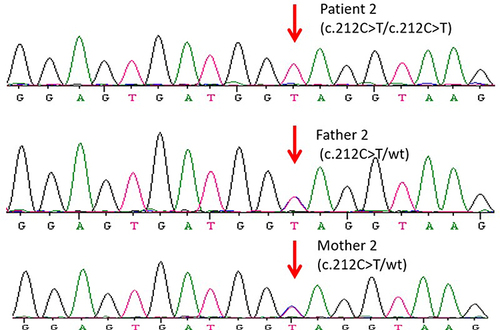
Table 1 Evaluation and Classification of the Three Genetic Variants Detected by WES
Figure 3 CMA testing result of Individual 1. A 24 Mb heterozygous deletion (LOH: loss of heterozygosity) in the 2p12.1p13.13 region was detected.
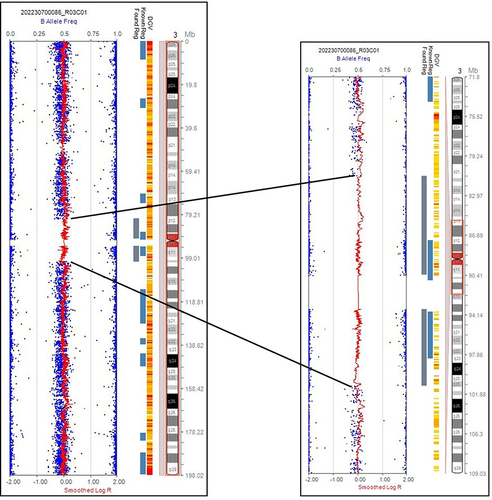
Table 2 Clinical Features, Laboratory results in Seven Individuals
