Figures & data
Table 1 Clinical Characteristics of Subjects
Figure 1 Expression of HOTAIR and changes in inflammation and APN. (A) RT-qPCR to test HOTAIR expression. (B–D) ELISA to analyze TNF-α, IL-10, and APN levels. All experiments were replicated thrice. *P < 0.05, **P < 0.01.
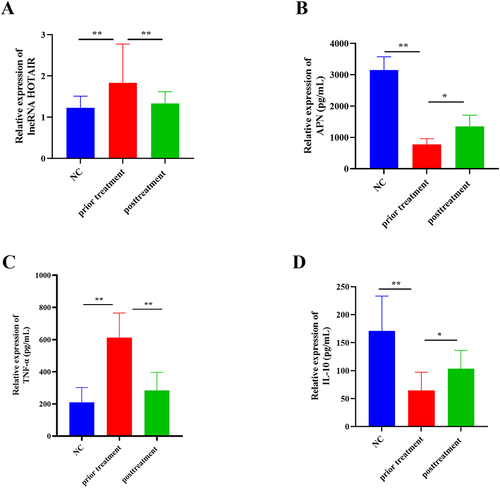
Table 2 Association Rule Analysis of JQP with Clinical Characteristics of OA Patients
Table 3 The Random Walk Evaluation Model of Laboratory Indicators of OA Patients Treated with JQP
Figure 2 Random walking analysis of JQP and clinical indicators.
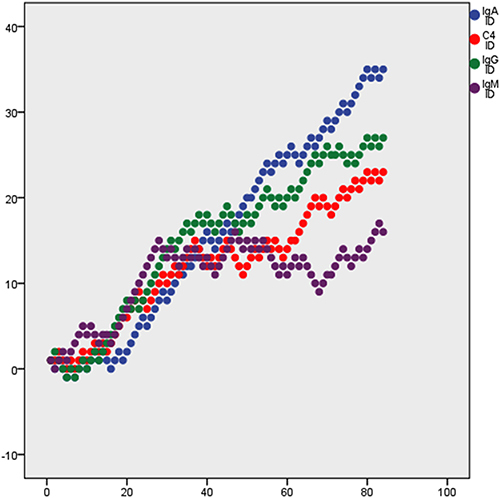
Table 4 Active Ingredient Information of JQP
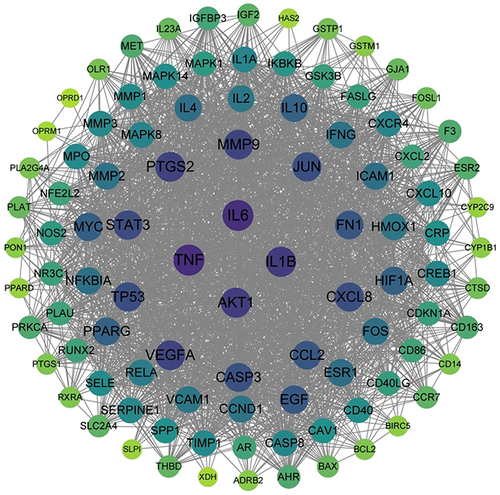
Figure 4 Network of Drug-Compound-Target of JQP. LGT: Tripterygium wilfordii Hook. f, WLX: Clematis chinensis Osbeck, ZZ: Gardenia jasminoides J. Ellis, HQIN: Scutellaria baicalensis Georgi, WG: Scolopendra subspinipes mutilans L. Koch, YYR: Coix lacryma-jobi L, TR: Juglans regia L, HQ: Astragalus mongholicus Bunge. Green arrow nodes: the eight Chinese herbal medicines in JQP; Orange diamond nodes: potential drug targets; pink octagon nodes: active compounds.
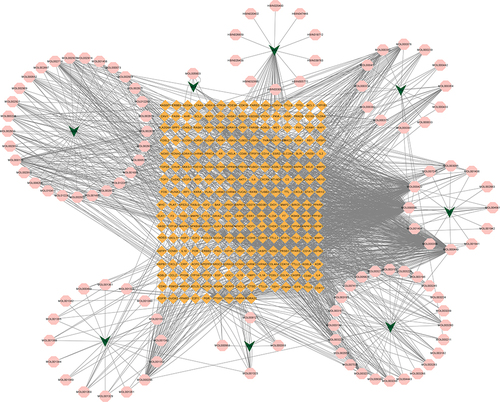
Table 5 The Core Active Ingredient of JQP for the Treatment of OA
Table 6 The Core Targets of JQP in OA Treatment
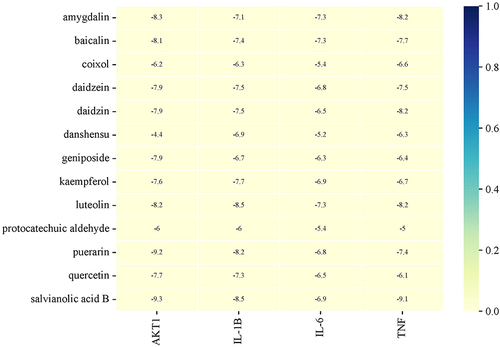
Figure 9 The representative docking complex of key targets and compounds of JQP in OA treatment, (A) AKT1 and puerarin; (B) AKT1 and salvianolic acid B; (C) TNF and salvianolic acid B.
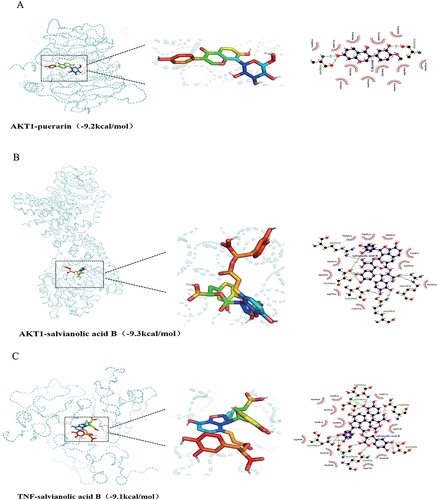
Figure 10 RT-qPCR to examine HOTAIR expression (A); ELISA to test IL-1β, IL-6, IL-4, IL-10, APN levels (B-F). *** P < 0.001; ** P < 0.01; * P < 0.05.
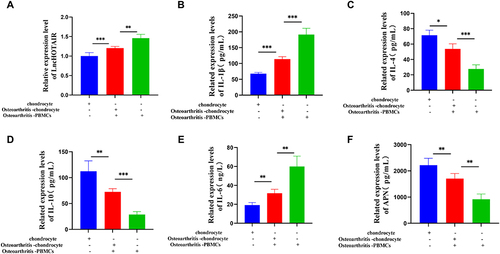
Figure 11 RT-qPCR to check HOTAIR expression (A and B); CCK-8 assay to analyze PC viability (C); WB to test phosphorylation of PI3K and AKT and levels of APN (D); ELISA to determine IL-1β, IL-6, IL-4, IL-10, and APN levels (E–H); Semi-quantitative levels of APN, p-PI3K, and p-AKT detected with immunofluorescence (I–K). *** P < 0.001; ** P < 0.01; * P < 0.05.
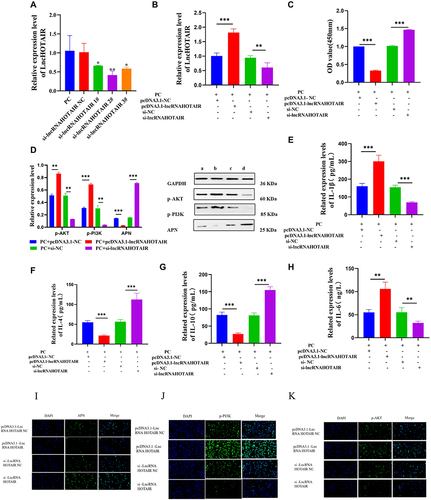
Figure 12 CCK-8 assay to examine CH viability (A); Phosphorylation of PI3K and AKT and level of APN measured with WB (B); IL-1β, IL-6, IL-4, IL-10, and APN levels tested with ELISA (C–F); Semi-quantitative levels of APN, p-PI3K, and p-AKT checked with immunofluorescence (G–I). *** P < 0.001; ** P < 0.01.
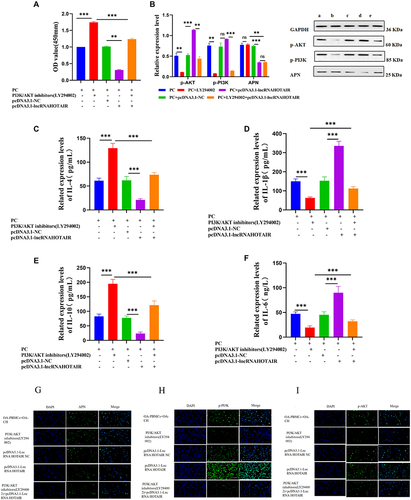
Figure 13 CH viability examined with CCK-8 assay (A); HOTAIR expression measured with PCR (B); Level of IL-1β, IL-6, IL-4, IL-10, and APN checked with ELISA (D-F); Phosphorylation of PI3K and AKT and level of APN tested with WB (G); Semi-quantitative levels of APN, p-PI3K, and p-AKT detected with immunofluorescence (H-J). ***P < 0.001; **P < 0.01; *P < 0.05.
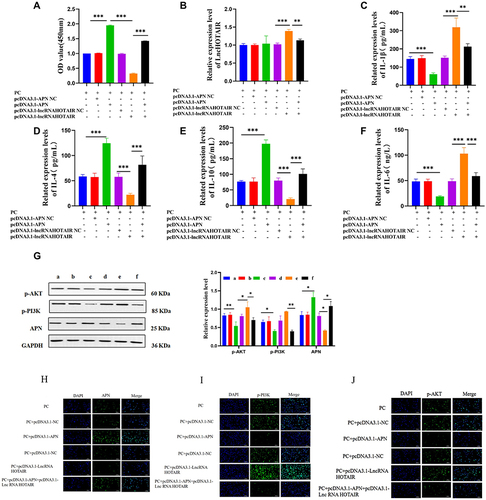
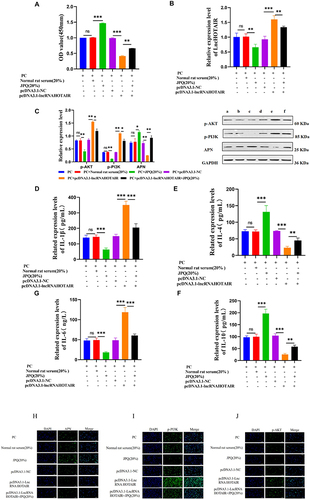
Figure 14 CH viability measured with CCK-8 assay (A); HOTAIR levels tested with PCR (B); Phosphorylation of PI3K and AKT and level of APN checked with WB (C); IL-1β, IL-6, IL-4, IL-10, and APN levels examined with ELISA (D-G); Semi-quantitative levels of APN, p-PI3K, and p-AKT determined with immunofluorescence (H-J). *** P < 0.001; ** P < 0.01; * P < 0.05.
Data Sharing Statement
The data that support the findings of this study are available from the information Center of Anhui Hospital of Traditional Chinese Medicine obtains the electronic medical record information of effective OA inpatients from the SQL Server database, but restrictions apply to the availability of these data, which were used under license for the current study, and so are not publicly available. Data are, however, available from the authors upon reasonable request and with permission of the information Center.