Figures & data
Figure 1 Depiction of the strategy used to prepare antibiofilm CNMA-GNPs and the model transportation of nanoparticle into the EPS layer for payload release, and eradication of biofilm by CNMA-GNPs have shown.
Abbreviations: CNMA, cinnamaldehyde; EPS, extracellular polymeric substance; GNPs, gold nanoparticles; TEOS, tetraethyl orthosilicate.
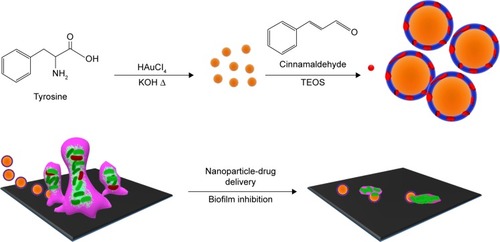
Figure 2 Physicochemical characteristics of the surface modified gold nanoparticles conjugated with or without cinnamaldehyde (CNMA): free CNMA (green), GNPs (black), Si-GNPs (red) and CNMA-GNPs (blue). (A) UV-Visible absorbance spectra, (B) FT-IR spectra, and (C) transmission electron micrographs (showing the surface coating on GNPs) of nano-dispersions.
Abbreviations: CNMA, cinnamaldehyde; GNP, gold nanoparticle; Si, silica.
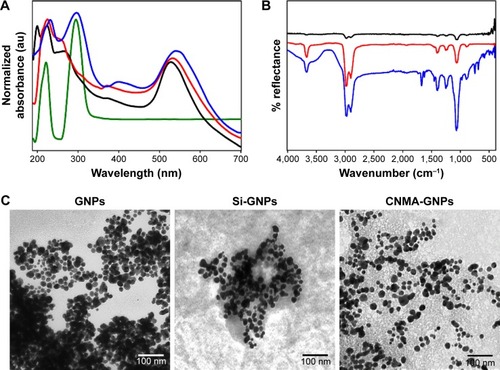
Figure 3 Effects of nano-dispersions on biofilm formation. The effects of different concentrations of (A) CNMA, (B) Si-GNPs and (C) CNMA-GNPs, on biofilm formation by E. coli O157:H7, P. aeruginosa, MSSA 6538, and MRSA were examined after incubation for 24 h in 96-well plates without shaking. At least three independent experiments were conducted. (D) Representative 3D projection confocal laser scanning microscopy images of E. coli O157:H7, P. aeruginosa, MSSA 6538, and MRSA biofilms after treatment with free CNMA, Si-GNPs, and CNMA-GNPs or none (control). *P<0.05 versus the control.
Abbreviations: CNMA, cinnamaldehyde; GNP, gold nanoparticle; MRSA, methicillin-resistant Staphylococcus aureus; MSSA, methicillin-sensitive Staphylococcus aureus; Si, silica.
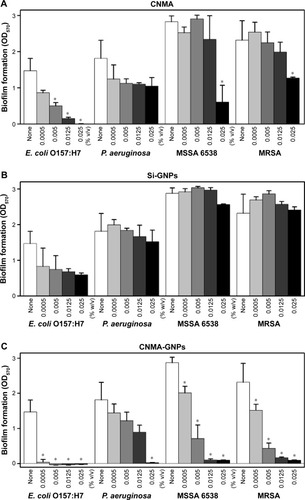
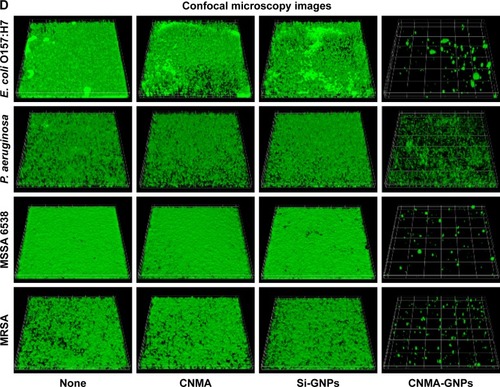
Figure 4 Morphologies of untreated and treated bacteria as determined by TEM. Low and high magnification images showing the ultrastructural changes induced in MSSA 6538 treated with nothing (A and A′), Si-GNPs (B and B′), or CNMA-GNPs (C and C′). Yellow arrow indicates nanocomposites and red arrow indicates cell damages. Elemental spectrum of the CNMA-GNPs treated bacteria depicting the presence of Au and Silica (D).
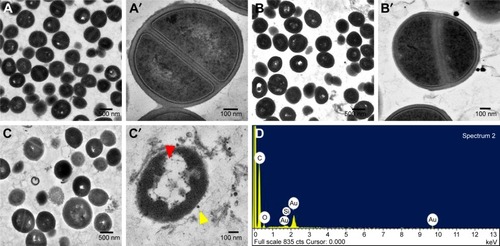
Figure 5 Kaplan–Meier survival curves for nematodes infected with S. aureus. Results of killing assays of C. elegans strain fer-15; fem-1 infected with S. aureus and fed with Si-GNPs or CNMAGNPs. Non-treated infected nematodes were considered as none. The percentage survivals shown represent the results of three independent experiments (n=60) performed. E. coli OP50 was the common food source and used as the control strain.
Abbreviations: CNMA, cinnamaldehyde; GNP, gold nanoparticle; Si, silica.
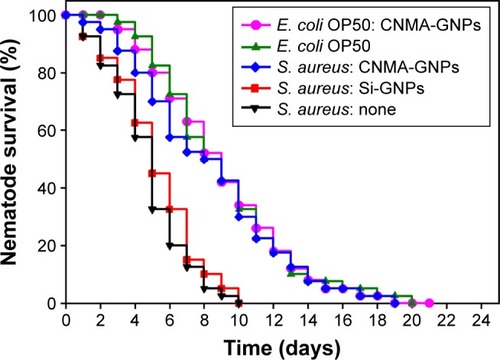
Figure 6 Microscopic study of nanoparticle internalizations by C. elegans. Phase contrast and dark field microscopy images of C. elegans nematodes; (A) Non-treated nematode (a- whole body; b- tail and c- head), (B) CNMA-GNP treated worms co-incubated with MSSA 6538 (a- whole body; b- tail and c- head of nematode) and (C) high magnification tail images (a- phase contrast, b- dark field). Arrows show individual and aggregated CNMA-GNPs.
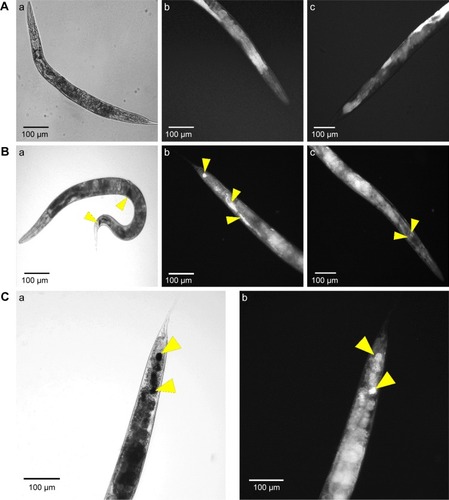
Figure S1 X-ray diffraction patterns of GNPs (black), Si-GNPs (red) and CNMA-GNPs (blue).
Abbreviations: CNMA, cinnamaldehyde; GNP, gold nanoparticle; Si, silica.
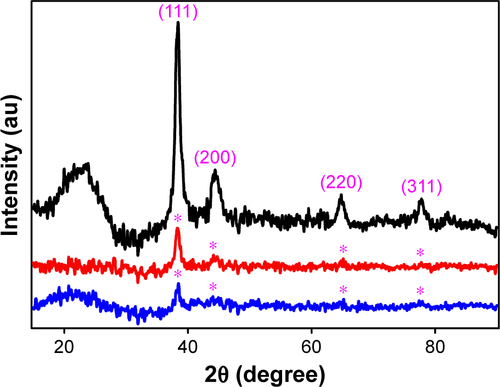
Figure S2 High resolution region XP spectra of CNMA-GNPs. (A) Survey spectrum, (B) C 1s, (C) O 1s, (D) N1s, (E) Si 2p, and (F) Au0 4f7/2 and 4f5/2, respectively. Black lines indicate experimental results, and colored lines representing different chemical states.
Abbreviations: CNMA, cinnamaldehyde; GNP, gold nanoparticle; Si, silica.
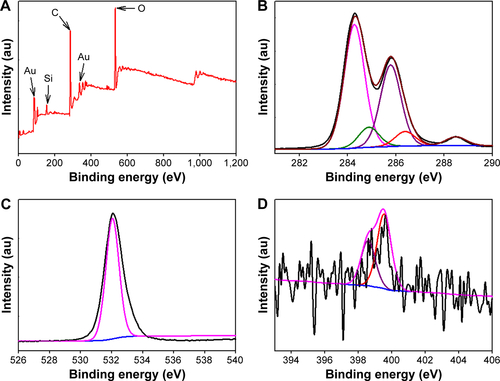
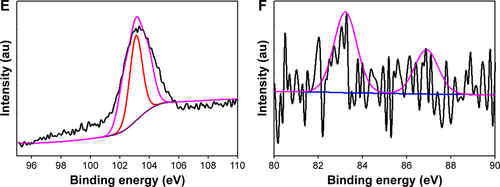
Figure S3 In vitro release profile of CNMA from CNMA-GNPs in different pH buffered saline at 37°C.
Abbreviations: CNMA, cinnamaldehyde; GNP, gold nanoparticle.
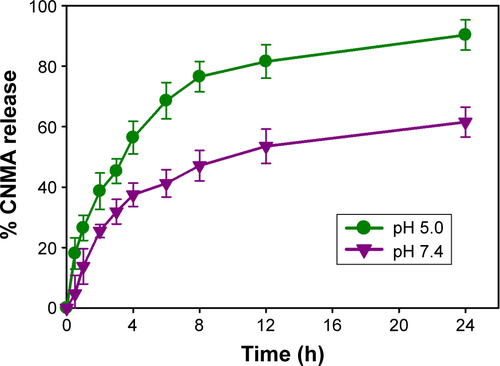
Figure S4 Planktonic cell growths of E. coli O157:H7, P. aeruginosa, S. aureus MSSA 6538 and S. aureus MRSA in the presence of (A) CNMA, (B) Si-GNPs and (C) CNMA-GNPs, measured at OD620.
Abbreviations: CNMA, cinnamaldehyde; GNP, gold nanoparticle; MRSA, methicillin-resistant Staphylococcus aureus; MSSA, methicillin-sensitive Staphylococcus aureus.
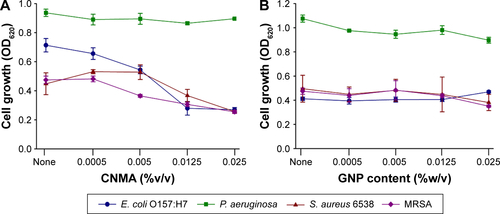
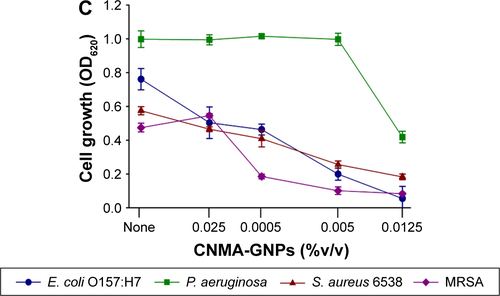
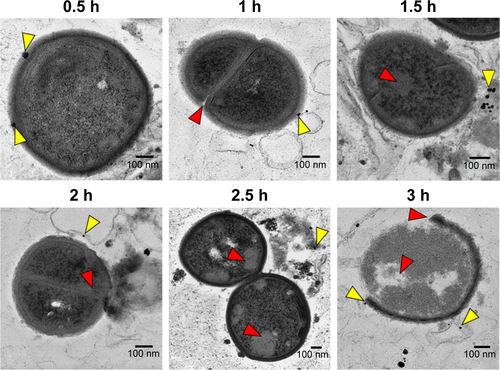
Figure S5 TEM findings and the mechanism of cell damage. Thin section photograph of a MSSA 6538 bacterium treated with CNMA-GNPs for 0.5 to 3 h. High magnification images showed the time-dependent penetration and bactericidal effect of CNMA-GNPs. The yellow arrows indicate nanoparticles binding with cells and the red arrows show the severity of cell damage.
Abbreviations: CNMA, cinnamaldehyde; GNP, gold nanoparticle; MSSA, methicillin-sensitive Staphylococcus aureus; TEM, transmission electron microscopy.