Figures & data
Table 1 Primer sequence of target gene for rats
Figure 1 CUS caused depression-like behavior in rats.
Abbreviations: CUS, chronic unpredictable stress; SPT, sucrose preference test; OFT, open field test; FST, forced swimming test; CT, computed tomography; HE, hematoxylin and eosin; BMSCs, bone marrow mesenchymal stem cells.
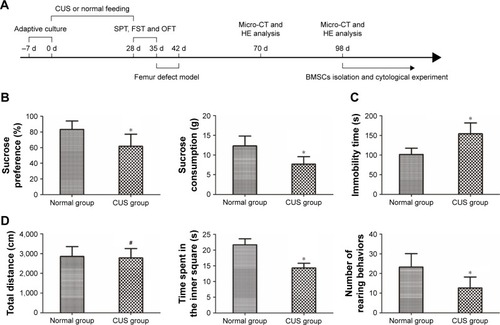
Figure 2 Depression attenuated bone healing in a rat femur defect model.
Abbreviations: CUS, chronic unpredictable stress; CT, computed tomography; BV/TV, bone volume/trabecular volume ratio; Tb.N, trabecular number; Tb.Th, trabecular thickness; Tb.Sp, trabecular separation; VOI, volume of interest; HE, hematoxylin and eosin.
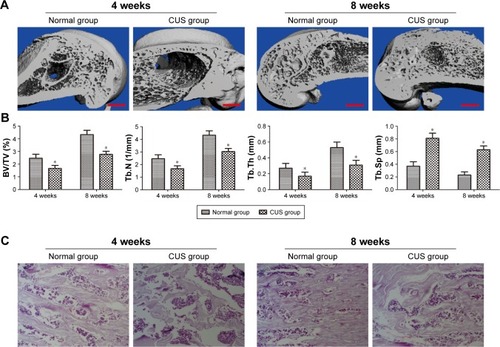
Figure 3 BMSCs derived from depressive rats had low osteogenic potential and autophagic level.
Abbreviations: ALP, alkaline phosphatase; ARS, Alizarin red S; CUS, chronic unpredictable stress; BMSCs, bone marrow mesenchymal stem cells; qPCR, quantitative polymerase chain reaction; TEM, transmission electron microscopy.
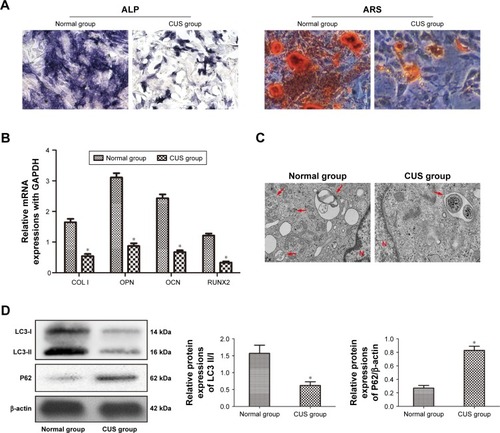
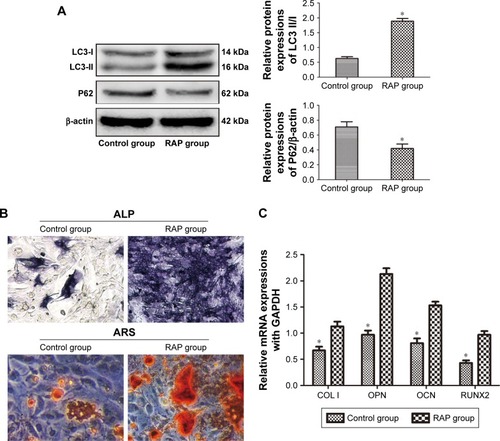
Figure 4 Autophagy activation promoted osteogenic differentiation of depressive BMSCs.
Abbreviations: RAP, rapamycin; ALP, alkaline phosphatase; ARS, Alizarin red S; BMSCs, bone marrow mesenchymal stem cells; qPCR, quantitative polymerase chain reaction.