Figures & data
Table 1 Correlation of ARHGAP30 expression with patient characteristics
Figure 1 ARHGAP30 is markedly downregulated in lung cancer tissues and cell lines.
***P<0.001.
Abbreviation: TCGA, The Cancer Genome Atlas.
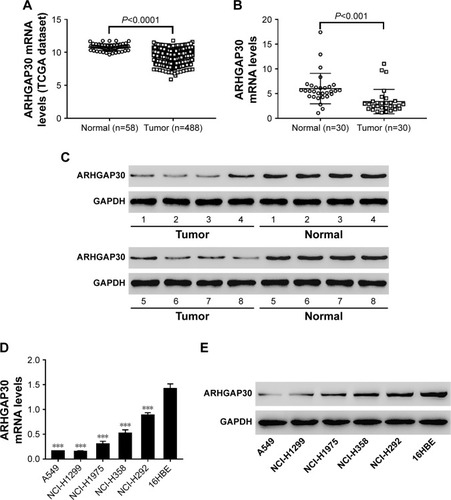
Figure 2 ARHGAP30 inhibited proliferation of lung cancer cells.
Abbreviations: ARH OE, ARHGAP30-overexpressing lentivirus; CCK-8, Cell Counting Kit-8; h, hours; NS, no significant difference; qRT-PCR, quantitative real-time PCR; WT, wild-type.
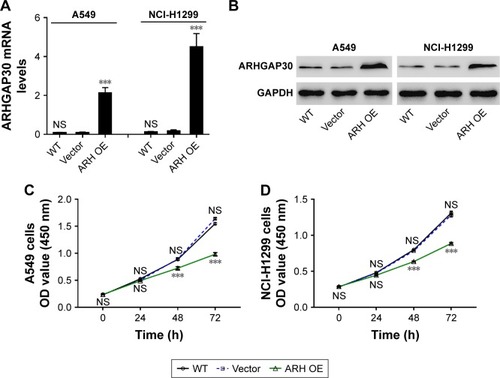
Figure 3 ARHGAP30 suppressed lung cell migration and invasion.
Abbreviations: ARH OE, ARHGAP30-overexpressing lentivirus; NS, no significant difference; qRT-PCR, quantitative real-time PCR; WT, wild-type.
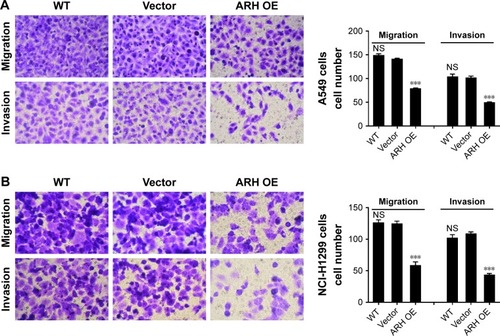
Figure 4 ARHGAP30 repressed the Wnt signaling pathway in lung cancer cells.
Abbreviations: ARH OE, ARHGAP30-overexpressing lentivirus; GSEA, gene set enrichment analysis; NES, normalized enrichment score; NS, no significant difference; qRT-PCR, quantitative real-time PCR; TCGA, The Cancer Genome Atlas.
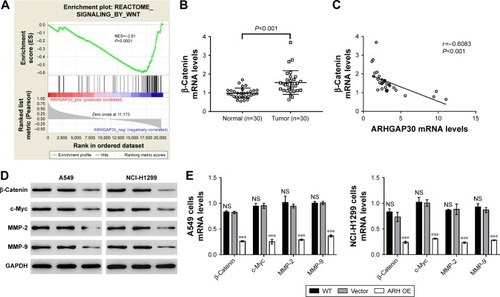
Figure 5 The Wnt signaling pathway mediated the effects of ARHGAP30 on the proliferation, migration, and invasion of lung cancer cells.
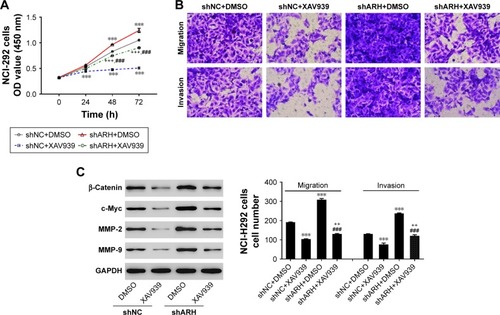
Figure S1 ARHGAP30 inhibited nuclear translocation of β-catenin.
Abbreviation: ARH OE, ARHGAP30-overexpressing lentivirus.
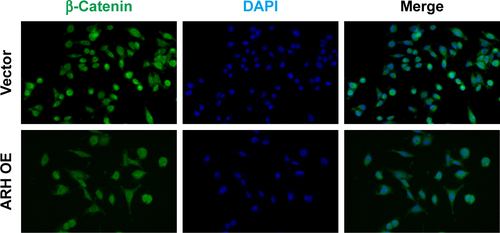
Figure S2 Downregulated ARHGAP30 expression in NCI-H292 cells.
Abbreviations: NS, no significant difference; qRT-PCR, quantitative real-time PCR; shNC, control shRNA; shRNA, short hairpin RNA.
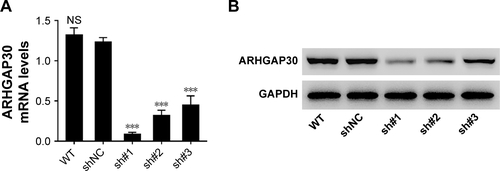
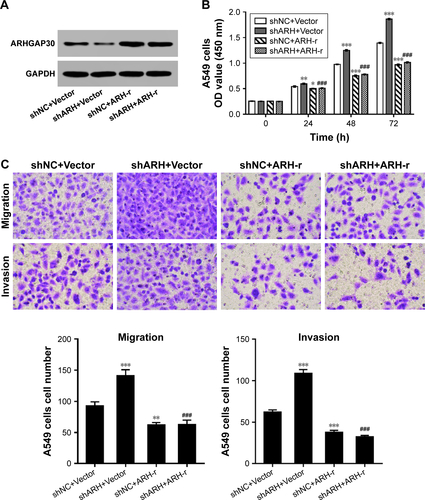
Figure S3 RNAi-resistant ARHGAP30 rescues the effects of ARHGAP30 knockdown.
Abbreviations: ARH-r, ARHGAP30 lentivirus; CCK-8, Cell Counting Kit-8; h, hours; shARH, ARHGAP30 shRNA; shNC, control shRNA; shRNA, short hairpin RNA.