Figures & data
Table 1 Primers Used in RT-qPCR
Figure 1 Profiles of HOTAIR and miR-34a expression in CSCs of PDAC.
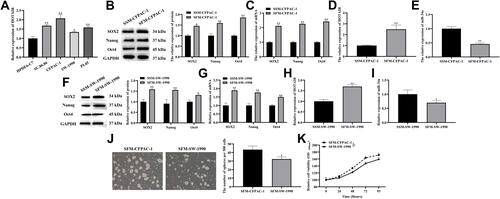
Figure 2 Knocking down HOTAIR inhibited CSCs-like properties, invasion and migration in PDAC.
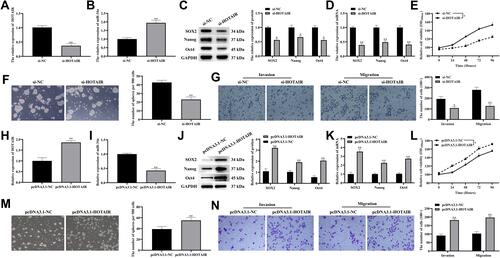
Figure 3 Overexpressing miR-34a inhibited CSCs-like properties, invasion and migration of PDAC.
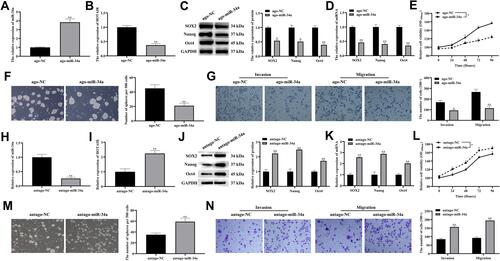
Figure 4 HOTAIR directly targets miR-34a.
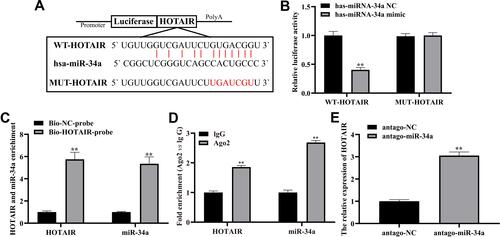
Figure 5 HOTAIR promoted CSCs properties, invasion and migration by interacting with miR-34a to activate the JAK2/STAT3 pathway in PDAC.
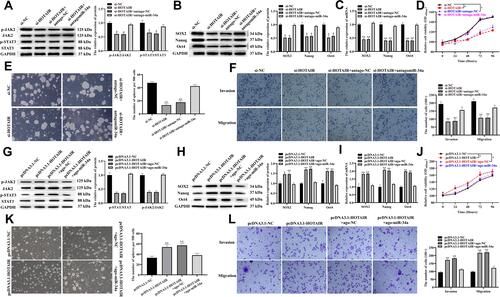
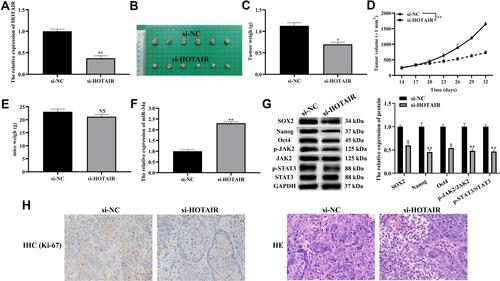
Figure 6 Knocking down HOTAIR inhibited tumorigenicity of CFPAC-1 cells.