Figures & data
Table 1.
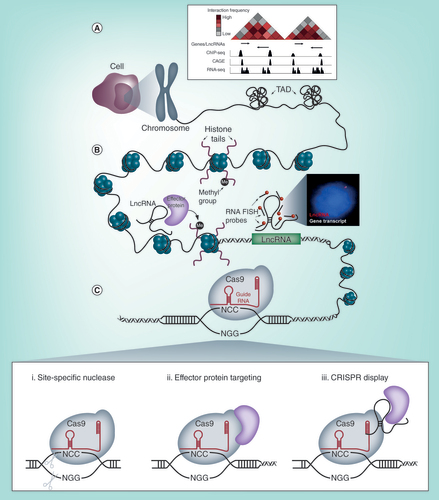
Summary of the emerging methods used to study long noncoding RNA biology.