Figures & data
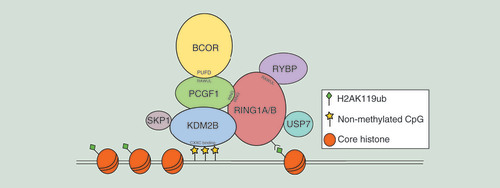
Table 1. PRC 1.1 complex components.
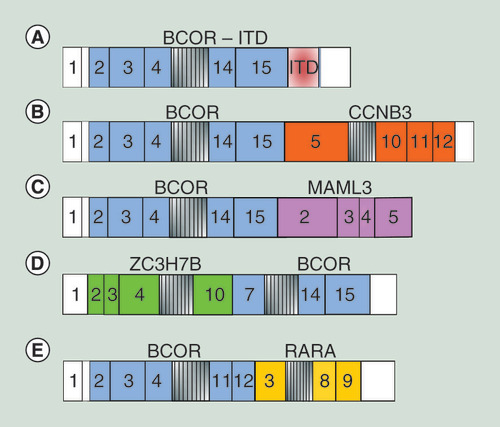
Table 2.
BCOR mutations in different tumor hystotypes.