Figures & data
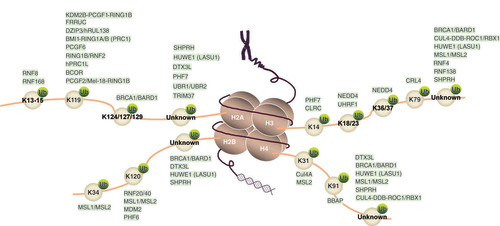
Table 1.
RING finger proteins ubiquitinate different lysine sites of core histones.
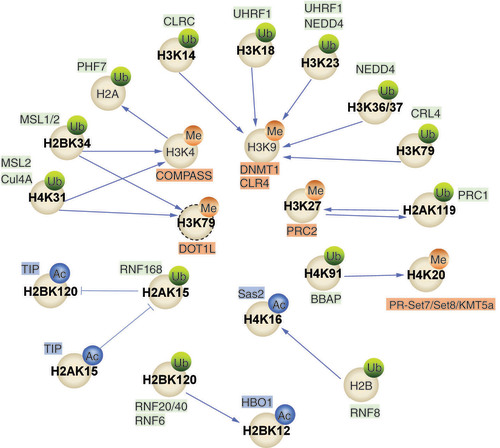
Table 2.
RING finger proteins mediate the crosstalk between histone ubiquitination and histone methylation; the relationship between the interactions is also shown.
Table 3.
RING finger proteins that mediate the crosstalk between histone ubiquitination and histone acetylation; the relationship between the interactions is also listed.