Figures & data
Table I. Pluronic information.
Figure 1. Pluronic thermosensitising effects on DHD/K12/TRb cells in vitro after 20 min of Pluronic and hyperthermia (at 43°C) co-exposure. Pluronic concentrations are 2.5 mg/mL. Data presented as Mean ± SD (standard deviation, n = 3). *Significant difference compared to Pluronic only; †significant difference compared to untreated control (P < 0.05).
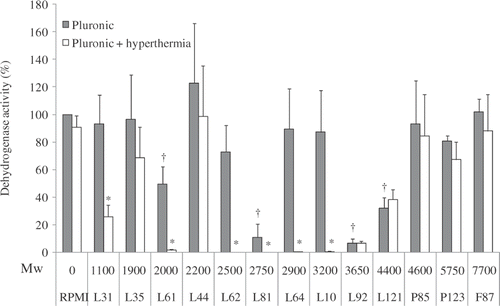
Figure 2. Dependence of Pluronic bioactivity on average molecular weight: (A) hydrophilic Pluronics; (B) hydrophobic Pluronics. Results are based on the dehydrogenase activity acquired with WST-1 assay. Data presented as Mean ± SD (standard deviation, n = 3).
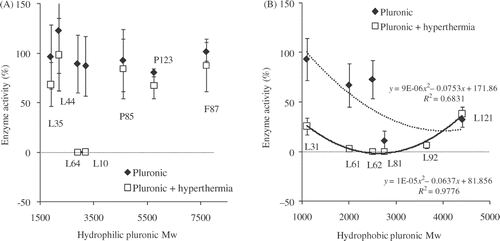
Figure 3. Caspase 3/7 activity. Data are normalised by the cell counts: (A) 2 h after treatment; (B) 24 h after treatment. Data presented as Mean ± SD (standard deviation, n = 4). *Significant difference versus hyperthermia only; ‡significant difference versus Pluronic only (P < 0.05).
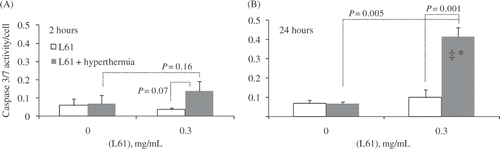
Figure 4. Change in tumour volume relative to size before treatment in response to radiofrequency ablation with or without Pluronic L61. Data presented as Mean ± SD (standard deviation, n = 17). Portions of the error bars were omitted for presentation clarity. *Indicates statistically significant difference (P < 0.05).
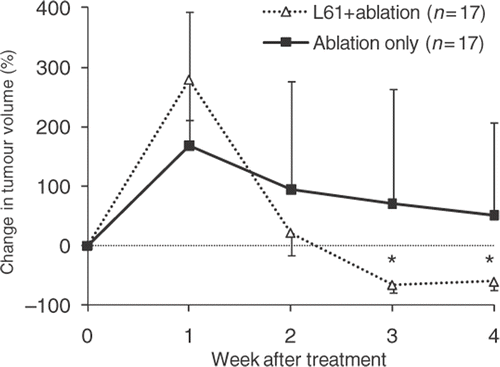
Figure 5. Proportion of tumours showing shrinkage of <100% in volume. Estimation was done with Kaplan-Meier survival methods and the statistics were acquired with the Cox proportional hazards regression model (P = 0.03).
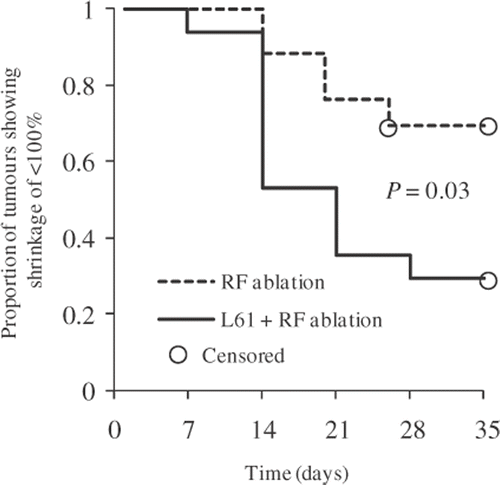
Figure 6. The four-group Pluronic copolymer grid based on Pluronic HLB and NPO adapted from a previously published model Citation[11]. Here the authors categorised the investigated Pluronic copolymers into four groups according to the HLB and NPO numbers. According to their results, Group II polymers were the most effective in modulating P-glycoprotein, while polymers from Group I and III showed little effect.
![Figure 6. The four-group Pluronic copolymer grid based on Pluronic HLB and NPO adapted from a previously published model Citation[11]. Here the authors categorised the investigated Pluronic copolymers into four groups according to the HLB and NPO numbers. According to their results, Group II polymers were the most effective in modulating P-glycoprotein, while polymers from Group I and III showed little effect.](/cms/asset/f995327b-4ded-4078-86db-d6cdac742351/ihyt_a_599828_f0006_b.gif)