Figures & data
Table I. Primer details for target and reference genes.
Table II. Networks identified of genes differentially expressed in male adult mice after neonatal infection (bold indicates up-regulation).
Figure 1. Network 1 identified by IPA showing differentially expressed genes in male adult hippocampus after neonatal respiratory C. muridarum infection. Red shading indicates up-regulation of the gene. No color indicates genes that form the network and have no change in expression in the present study. Fourteen genes were differentially expressed and associated with tissue morphology, cellular development, and connective tissue development and function. For example, 11β-HSD1 gene expression was altered following male neonatal infection and was indirectly regulated by IL-4. For full names of genes involved in the network, see .
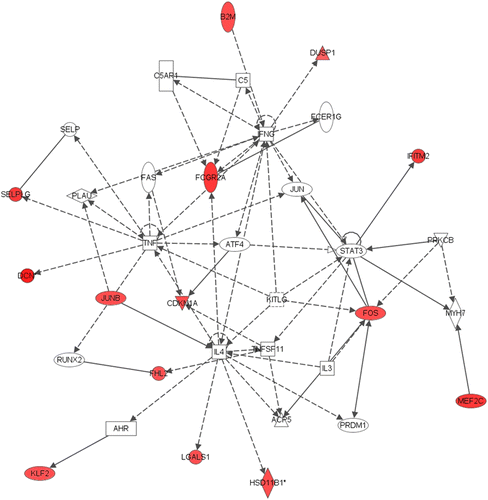
Table III. Networks identified of genes differentially expressed in female adult mice after neonatal infection (bold indicates up-regulation; italic indicates down-regulation).
Table IV. Definitions of abbreviations used in figures and tables.
Figure 2. Network 1 identified by IPA showing differentially expressed genes in female adult hippocampus after neonatal respiratory C. muridarum infection. Red shading indicates up-regulation of the gene. Gray shading indicates down-regulation. No color indicates genes that form the network and have no change in expression in the present study. Thirteen genes were differentially expressed after infection and were indirectly involved with cytokine regulation. For example, HLA DQB2 was increased following infection and identified to indirectly regulate IL-6 and IL-10. For full names of genes involved in the network, see .
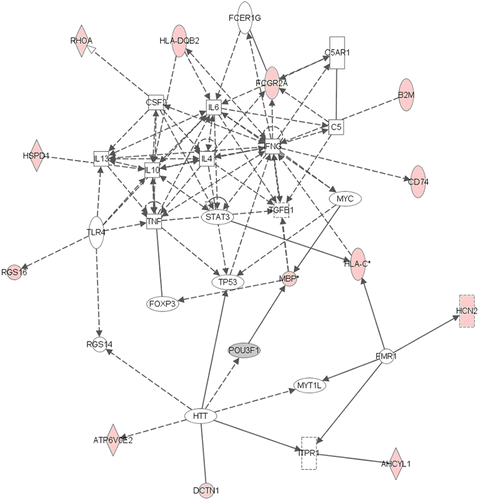
Table V. Direction of change in mRNA in microarray (fold change) and qRT PCR (relative abundance) with pairwise comparison across treatment for qRT PCR.