Figures & data
Figure 2. I338/I332 intensity ratio calculated from pyrene excitation spectra and I1/I3 intensity ratio calculated from pyrene emission spectra in water, as a function of the logarithm of copolymer concentration.
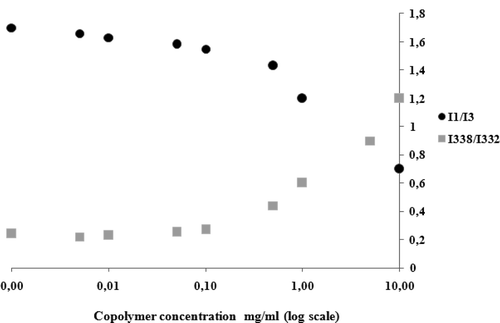
Figure 3. Contour plot of the NOESY experiment on PHEA-EDA-Sq17-PS80 copolymer micelles recorded with an experimental mixing time of 350 ms.
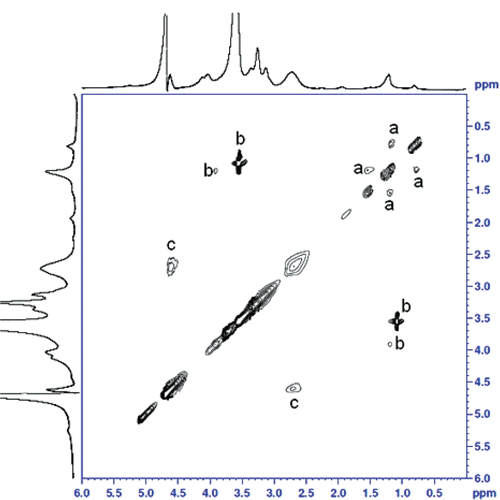
Table 1. Mean diameter, polydispersity index (PDI) and ζ potential of unloaded and Riv-loaded PHEA-EDA-Sq17-PS80 micelles.
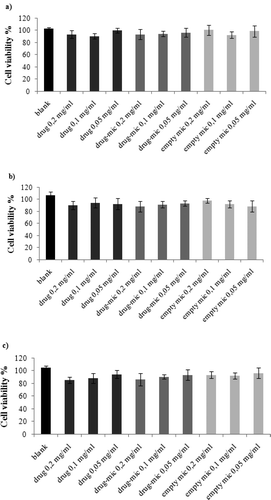
Figure 4. Cell viability of Neuro2a cells incubated for 6 h (a), 24 h (b) and 48 h (c) with free Riv (drug), Riv-loaded micelles (drug-mic) and unloaded micelles (empty mic). Each bar represents the mean of three sample ± SD.