Figures & data
Figure 1. Schematic representation of carboxymethylation (a) and propylation (b) of Hb through reductive alkylation chemistry.
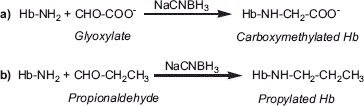
Table 1. Tetramer-Dimer Dissociation Constants of Modifed Hbs.
Figure 2. Size exclusion chromatography analysis of the hexaPEGylated Hbs. hexaPEGylated αα-Hb (1), hexaPEGylated Hb (2), hexaPEGylated [cm-Val-1(α)]2-Hb (3), hexaPEGylated [cm-Val-1(β)]2-Hb (4), were loaded on two HR10/30 Superose 12 columns and eluted with PBS, pH 7.4 at a flow rate of 0.5 ml/min.
![Figure 2. Size exclusion chromatography analysis of the hexaPEGylated Hbs. hexaPEGylated αα-Hb (1), hexaPEGylated Hb (2), hexaPEGylated [cm-Val-1(α)]2-Hb (3), hexaPEGylated [cm-Val-1(β)]2-Hb (4), were loaded on two HR10/30 Superose 12 columns and eluted with PBS, pH 7.4 at a flow rate of 0.5 ml/min.](/cms/asset/935b700d-10d2-4d63-b06d-57e56ad1c0cb/ianb19_a_501756_f0002_b.gif)
Figure 3. Size exclusion chromatography analysis of the PEGylated proteins. HbA (a), [Propyl-Val-1(α)]2-Hb (b), αα-fumaryl Hb (c), the hexaPEGylated Hb (d), the hexaPEGylated [Propyl-Val-1(α)]2-Hb (e), and the hexaPEGylated αα-fumaryl Hb (f) were loaded on two HR10/30 Superose 12 columns (1 × 31 cm2) and eluted with PBS, pH 7.4 at a flow rate of 0.5 ml/min.
![Figure 3. Size exclusion chromatography analysis of the PEGylated proteins. HbA (a), [Propyl-Val-1(α)]2-Hb (b), αα-fumaryl Hb (c), the hexaPEGylated Hb (d), the hexaPEGylated [Propyl-Val-1(α)]2-Hb (e), and the hexaPEGylated αα-fumaryl Hb (f) were loaded on two HR10/30 Superose 12 columns (1 × 31 cm2) and eluted with PBS, pH 7.4 at a flow rate of 0.5 ml/min.](/cms/asset/49682263-764a-403b-8b76-34cf528e7b68/ianb19_a_501756_f0003_b.gif)
Figure 4. RP-HPLC analysis of the PEGylated proteins. HPLC analysis of HbA (a), [Propyl-Val-1(α)]2-Hb (b), the hexaPEGylated Hb (c), and the hexaPEGylated [Propyl-al-1(α)]2-Hb (d) was carried out on a Vydac C4 column (4.6 × 250 mm2). The column was eluted with a linear gradient of 35–50% acetonitrile containing 0.1% TFA in 100 min and 50–70% acetonitrile containing 0.1% TFA in 30 min at a flow rate of 1.0 ml/min.
![Figure 4. RP-HPLC analysis of the PEGylated proteins. HPLC analysis of HbA (a), [Propyl-Val-1(α)]2-Hb (b), the hexaPEGylated Hb (c), and the hexaPEGylated [Propyl-al-1(α)]2-Hb (d) was carried out on a Vydac C4 column (4.6 × 250 mm2). The column was eluted with a linear gradient of 35–50% acetonitrile containing 0.1% TFA in 100 min and 50–70% acetonitrile containing 0.1% TFA in 30 min at a flow rate of 1.0 ml/min.](/cms/asset/8d65457d-2843-481e-a05b-6593dc35cd57/ianb19_a_501756_f0004_b.gif)
Figure 5. The tetramer-dimer dissociation constant (Kd) of the PEGylated proteins. HbA (a), the hexaPEGylated [Propyl-Val-1(α)]2-Hb (b), the hexaPEGylated Hb prepared using extension arm facilitated PEGylation (c), and the diPEGylated Hb with PEGylation at Val-1(β) (d) were diluted to a series of Hb concentrations and subjected to two Superose 12 columns (1 × 31 cm2) in series for the Kd measurement.
![Figure 5. The tetramer-dimer dissociation constant (Kd) of the PEGylated proteins. HbA (a), the hexaPEGylated [Propyl-Val-1(α)]2-Hb (b), the hexaPEGylated Hb prepared using extension arm facilitated PEGylation (c), and the diPEGylated Hb with PEGylation at Val-1(β) (d) were diluted to a series of Hb concentrations and subjected to two Superose 12 columns (1 × 31 cm2) in series for the Kd measurement.](/cms/asset/955aa1f0-a72f-4f0e-821e-6a3fb431acf5/ianb19_a_501756_f0005_b.gif)
Table 2. The oxygen affinity of the HexaPEGylated [Propyl-Val-1(α)]2-Hb Sample.
Figure 6. Molecular models of [Propyl-PEG5K-Val-1(α)]2-Hb and [Propyl-PEG5K-Val-1(β)]2-Hb. The α- and the β-globin chains are shown in red and blue, respectively. The models were generated as described under Experimental Procedures.
![Figure 6. Molecular models of [Propyl-PEG5K-Val-1(α)]2-Hb and [Propyl-PEG5K-Val-1(β)]2-Hb. The α- and the β-globin chains are shown in red and blue, respectively. The models were generated as described under Experimental Procedures.](/cms/asset/92964cbd-8808-4d60-ba53-a1ec0626c4ad/ianb19_a_501756_f0006_b.jpg)
Table 3. Loss of Accessible Surface Area on PEGylation of Hb.
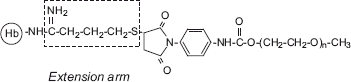
Figure 7. Schematic representation of the extension arms sandwiched between the PEG-chains and side chains of Hb in the EAF-PEGylated Hb. Note the extension arm between the amino group and succinimidyl phenyl PEG chains, which helps to attenuate the influence of the PEG-chains on the interdimeric interactions (central cavity interactions) of Hb tetramer.