Figures & data
Figure 1 . Structure of the caffeic acid derivative (7′Z)-3-O-(3, 4-dihydroxyphenylethenyl)-caffeic acid (CADP).
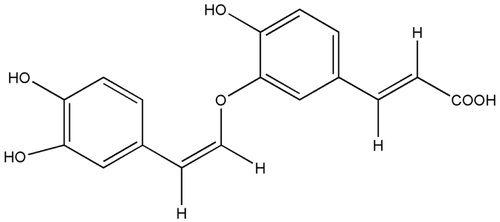
Figure 2 . Protective effect of (7′Z)-3-O-(3, 4-dihydroxyphenyl-ethenyl)-caffeic acid (CADP) against H2O2-induced PC12 cell injury. Cells were incubated with 200 μM H2O2 for 4 h for MTT assay. CADP was added to the culture 24 h before H2O2 addition. The data are presented as means ± SEM (n = 6). #p < 0.05 and ##p < 0.01 compared with H2O2 group.
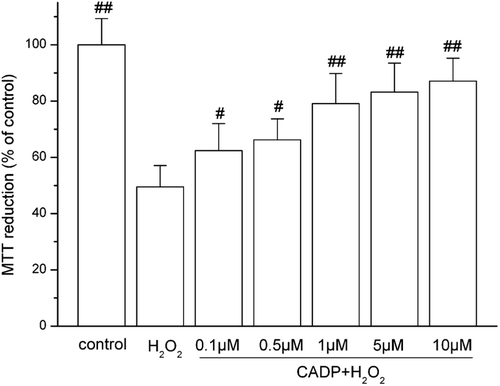
Figure 3 . Effects of (7′Z)-3-O-(3, 4-dihydroxyphenylethenyl)-caffeic acid (CADP) on the spatial learning and memory of mice in the Morris water maze test. (A) Mean latency in the hidden platform test during five consecutive days training. (B) Swimming distance moved in the hidden platform test during five consecutive days training. (C) The number of crossings over the exact location of the former platform. (D) Time spent in the target quadrant. All values are expressed as means ± SEM (n = 6–7). #p < 0.05 and ##p < 0.01 compared with the d-galactose (d-gal) model group.
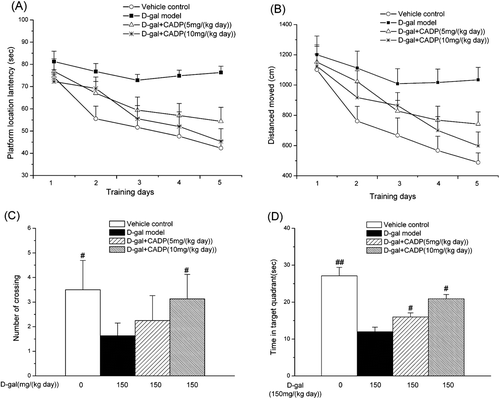
Figure 4 . The step-down avoidance test of all groups of mice. (A) The average step-down latency; (B) The average number of errors in 3 min. Data are expressed as means ± SEM (n = 6–7). #p < 0.05 and ##p < 0.01 compared with the d-galactose (d-gal) model group.
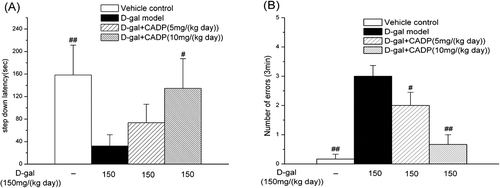
Table 1. Effect of CADP on MDA levels and activities of SOD, Gpx, and catalase in mouse brain.