Figures & data
Figure 1. Active sites of human BChECitation4. The enzyme active sites consist of at least five major binding sites: (1) an oxyanion hole (OAH); (2) an esteratic site (ES) or catalytic triad; (3) an anionic substrate binding site (AS); (4) an acyl binding site (ABS); and (5) a peripheral anionic binding site (PAS).
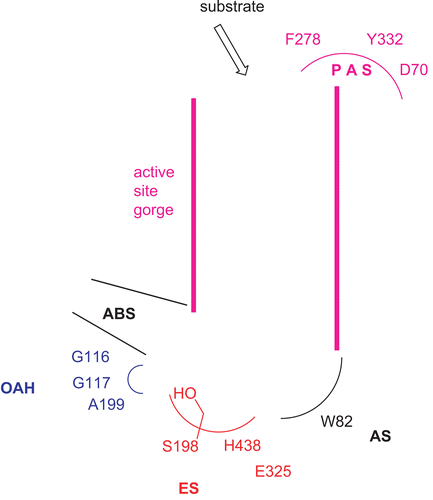
Figure 2. Structures of (R)-(+)-exo-, (S)-(–)-exo-, (R)-(+)-endo-, (S)-(–)-endo-2-norbornyl-N-n-butylcarbamates, rivastigmine, physostigmine, and carbaryl.
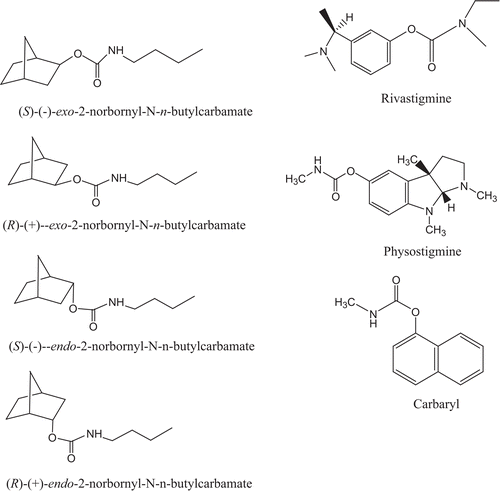
Figure 3. 19F-NMR spectra after the reaction of (A) (R)-(–)-exo-2-norborneol with S-(+)-α-methoxy-α-trifluoromethylphenylacetyl chlorideCitation39 in the presence of pyridine in CDCl3 and (B) (S)-(–)-exo-2-norborneol with (S)-(+)-α-methoxy-α-trifluoromethylphenylacetyl chloride in the presence of pyridine in CDCl3. For (A), −72.069−ppm was the fluorine chemical shift of unreactive (S)-(+)-α-methoxy-α-trifluoromethylphenylacetyl chloride. The peaks at −73.948 and −74.113 ppm were assigned to the fluorine chemical shifts of (2R)- and (2S)-exo-norbornyl-(S)-α-methoxy-α-trifluoromethylphenylacetates, respectively (). For (B), −72.089 ppm was the fluorine chemical shift of unreactive S-(+)-α-methoxy-α-trifluoromethylphenylacetyl chloride. The peaks at −73.965 and −74.130 ppm were assigned to the fluorine chemical shifts of (2R)- and (2S)-exo-norbornyl-(S)-α-methoxy-α-trifluoromethylphenylacetates, respectively ().
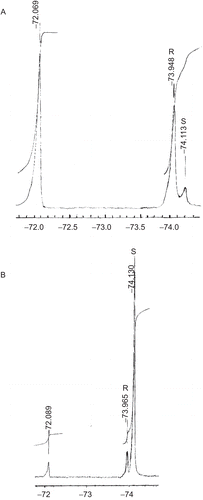
Table 1. Enantiomeric excess (%) and optical purity (%) for the kinetic resolution of racemic exo-2-norborneol () and endo-2-norborneol () by lipase in organic solvent.
Figure 4. Nonlinear least-squares curve fittings of (A) the kapp vs. (R)-(+)-exo-2-norbornyl-N-n-butylcarbamate concentration ([I]) plot and (B) the kapp vs. (R)-(+)-exo-2-norbornyl-N-n-butylcarbamate concentration ([I]) plot following Equation (1) for pseudo substrate inhibition of BChE. For , the parameters of the fit were k2 = 0.0030 ± 0.0001 s−1 and (1 + Km/[S])Ki = 22 ± 4 nM with R= 0.9963. After calculation, Ki = 11 ± 2 nM and ki = (270 ± 50) × 103 M− 1 s−1 (). For , the parameters of the fit were k2 = 0.00310 ± 0.00007 s−1 and (1 + Km/[S])Ki = 100 ± 4 nM with R= 0.9920. After calculation, Ki = 50 ± 2 nM and ki = (62 ± 3) × 103 M− 1 s−1 ().
![Figure 4. Nonlinear least-squares curve fittings of (A) the kapp vs. (R)-(+)-exo-2-norbornyl-N-n-butylcarbamate concentration ([I]) plot and (B) the kapp vs. (R)-(+)-exo-2-norbornyl-N-n-butylcarbamate concentration ([I]) plot following Equation (1) for pseudo substrate inhibition of BChE. For Figure 4A, the parameters of the fit were k2 = 0.0030 ± 0.0001 s−1 and (1 + Km/[S])Ki = 22 ± 4 nM with R= 0.9963. After calculation, Ki = 11 ± 2 nM and ki = (270 ± 50) × 103 M− 1 s−1 (Table 2). For Figure 4B, the parameters of the fit were k2 = 0.00310 ± 0.00007 s−1 and (1 + Km/[S])Ki = 100 ± 4 nM with R= 0.9920. After calculation, Ki = 50 ± 2 nM and ki = (62 ± 3) × 103 M− 1 s−1 (Table 2).](/cms/asset/5c7b4660-832b-42bc-90a0-e4bebe3ecd7d/ienz_a_388992_f0004_b.gif)
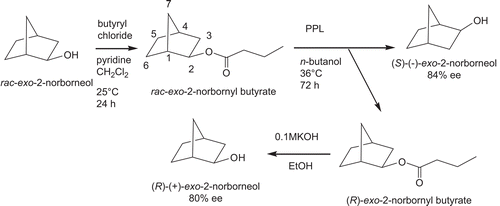
Scheme 1. Kinetic resolution of (R)-(+)- and (S)-(–)-exo-2-norborneols from lipase-catalyzed hydrolysis of racemic (±)-exo-2-norbornyl butyrate.
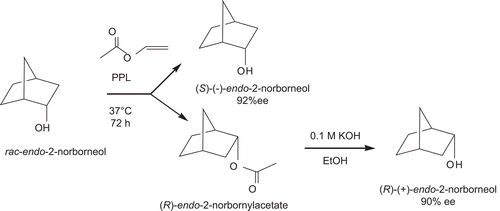
Scheme 2. Kinetic resolution of (R)-(+)- and (S)-(–)-endo-2-norborneols from lipase-catalyzed acetylation of racemic (±)-exo-2-norborneol with vinyl acetate.
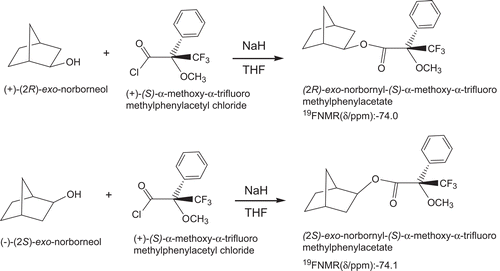
Scheme 3. Determination of enantiomeric excess and absolute configuration of (R)-(+)- and (S)-(–)-exo-2-norborneols by 19F-NMR spectra of their Mosher’s ester derivatives.
Scheme 4. Kinetic scheme for pseudo substrate inhibition of BChE by 2-norbornyl-N-n-butylcarbamate in the presence of substrate. E, enzyme; E-A, acyl enzyme; EI, enzyme–inhibitor Michaelis complex; E-I9, carbamyl enzyme; ES, enzyme–substrate Michaelis complex; I, pseudo substrate inhibitor; k2, carbamylation constant; k3, decarbamylation constant; k2S, formation rate constant of E-A; k3S, deacylation constant of E-A; Ki, inhibition constant; Km, Michaelis–Menten constant; P, product, 2-norborneol; P9, product, thiocholine; P99, product, butyrate; Q, product, butylcarbamic acid; S, substrate, BTCh.
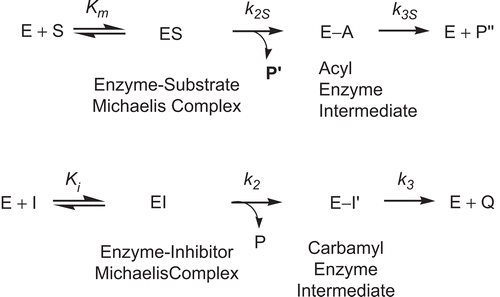
Table 2. The k2, Ki, and ki valuesa of BChE inhibitions by stereoisomers of 2-norbornyl carbamates.
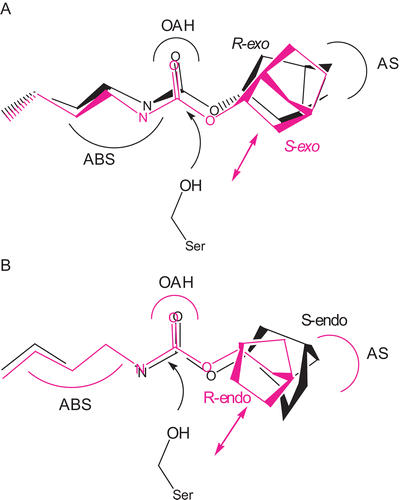
Figure 5. Superimposition of (A) (R)-(+)- and (S)-(–)-exo-2-norbornyl-N-n-butyl- carbamates and (B) (R)-(+)- and (S)-(–)-endo-2-norbornyl-N-n-butylcarbamates at their carbamyl moieties and fitting both enantiomers into the active site of BChE. For (A), an unfavorable repulsion between S-enantiomer and the active serine of the enzyme was observed. For (B), an unfavorable repulsion between R-enantiomer and the active serine of the enzyme was observed.