Figures & data
Table 1. Structures and observed experimental binding affinity of N-methylpiperazinylquinoxaline derivatives.a
Table 2. Physical meaning, average regression coefficients and the total incidences, and MLR-like coefficients from PLS model of descriptors identified from four-parameter CP-MLR models for the binding affinity of N-methylpiperazinylquinoxaline derivatives.
Table 3. Observed and modeled binding affinity of N-methylpiperazinylquinoxaline derivatives included in the training set.
Table 4. Observed, modelled, and residuala binding affinity of N-methylpiperazinylquinoxaline derivatives included in the test set.
Figure 1. Plot of observed versus calculated pKi values for the training (Δ) and test set (○) compounds. A, B, C, and D correspond, respectively, to four-parameter models (5), (6), (7), and (8) identified through CP-MLR. E corresponds to the MLR-like PLS equation from 18 CP-MLR-identified descriptors.
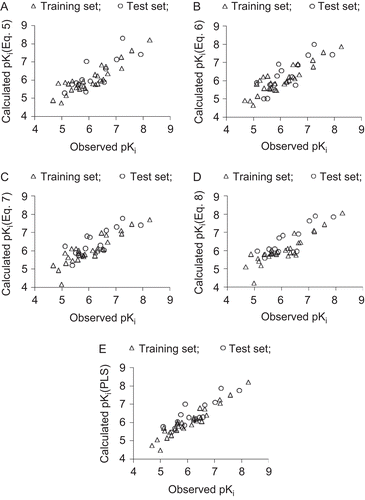
Figure 2. Plot of fraction contribution of MLR-like PLS coefficients (normalized) of the 18 descriptors () to the activity. The 10 most significant descriptors are identified by black-shaded lines. The numerals on bars are the order of the descriptors.
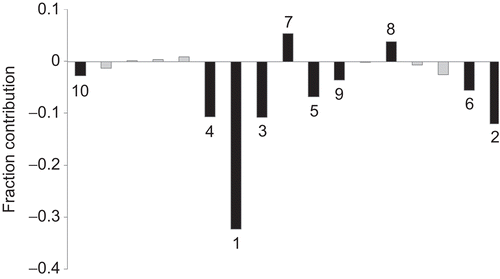
Table 5. Resultant models for the whole dataset (n = 44) in descriptors of training set models.
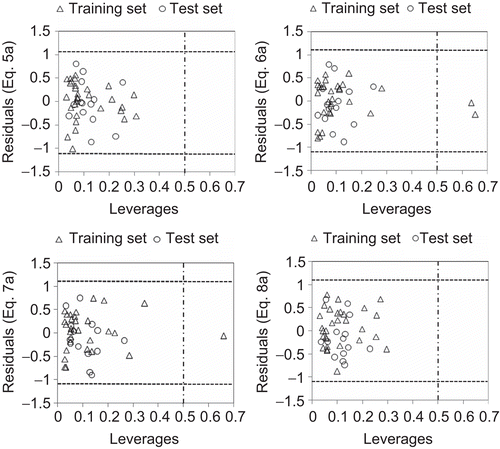
Figure 3. Williams plot for the training set and external prediction set for H4R binding affinity of N-methylpiperazinylquinoxaline derivatives.