Figures & data
Table 1. Structural features, observed and predicted ERβ IC50 values and ERα/β fold selectivity for 2-phenylquinoline scaffold.
Table 2. Structural features, observed and predicted ERβ IC50 values and ERα/β fold selectivity for tetrahydrofluorenone scaffold.
Table 3. Structural features, observed and predicted values for ERβ IC50 values and ERα/β fold selectivity for 3-hydroxy- 6H-benzo[c]chromen-6-one scaffold.
Table 4. Molecular descriptors selected for the study.
Table 5. Physicochemical descriptors that significantly influenced the 2-phenylquinoline derivatives binding at ER receptors.
Figure 1. Skeletal structure of estrogenic compounds. The moieties derivatised in each compound is indicated and referred in the , , and .
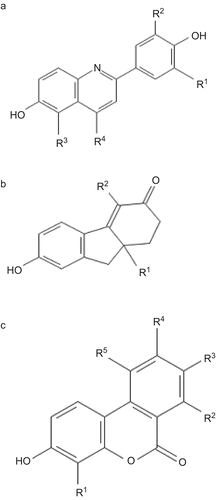
Table 6. QSAR statistics of the best multiple linear regression equations.
Figure 2. Plot of predicted vs. observed values of ERβ binding affinity and fold selectivity of the ligands against ER receptors. (A), (C) and (E) represent the ERβ binding affinity of phenylquinoline, tetrahydrofluorenone and 3-hydroxy 6H-benzo[c]chromen-6-one ligands respectively. (B), (D) and (F) represent the ERβ fold selectivity of phenylquinoline, tetrahydrofluorenone and 3-hydroxy 6H-benzo[c]chromen-6-one ligands, respectively.
![Figure 2. Plot of predicted vs. observed values of ERβ binding affinity and fold selectivity of the ligands against ER receptors. (A), (C) and (E) represent the ERβ binding affinity of phenylquinoline, tetrahydrofluorenone and 3-hydroxy 6H-benzo[c]chromen-6-one ligands respectively. (B), (D) and (F) represent the ERβ fold selectivity of phenylquinoline, tetrahydrofluorenone and 3-hydroxy 6H-benzo[c]chromen-6-one ligands, respectively.](/cms/asset/3df81295-0e67-4d90-a1be-b23d905be6db/ienz_a_566219_f0002_b.gif)
Figure 3. (A) Docked pose of most active compound 7 (green) and least active compound 21 (grey) in the phenylquinoline series with ERβ. Only key residues (pink), of the ERβ binding site are shown for simplicity. (B) Docked pose of most active compound 28 (green) and least active compound 7 (grey) in the tetrahydrofluorenone series with ERβ. Only key residues (pink), of the ERβ binding site are shown for simplicity. (C) Docked pose of most active compound 23 (green) and least active compound 7 (grey) in the tetrahydrofluorenone series with ERβ. Only key residues (pink), of the ERβ binding site are shown for simplicity.
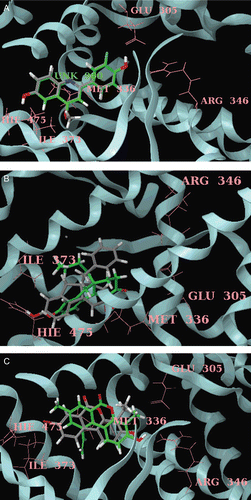
Table 7. Physicochemical descriptors that significantly influenced the tetrahydrofluorenone derivatives binding at ER receptors.
Table 8. Physicochemical descriptors that significantly influenced the 3-hydroxy 6H-benzo[c]chromen-6-one derivatives binding at ER receptors.