Figures & data
Figure 2. A snapshot picture taken from the simulation box: Protein (CA IX-dimer in yellow, inhibitor (inside the protein), water (as quick surface) and K+, Cl− ions (cyan and brown colored spheres).
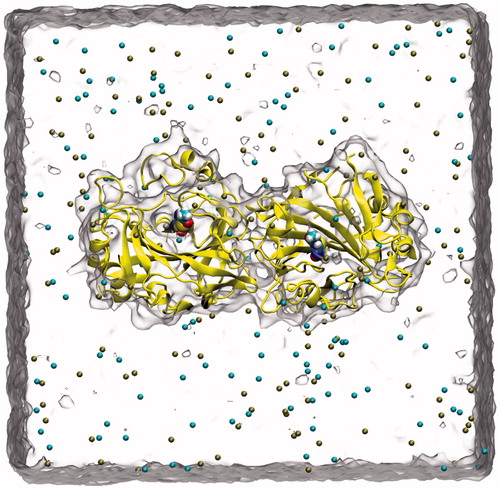
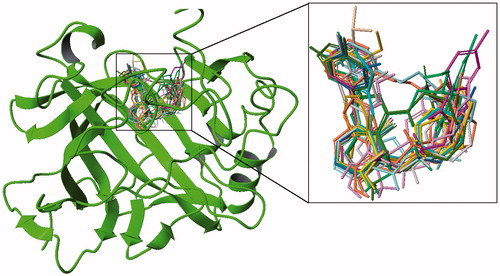
Figure 3. Cognate docking poses alignment with the co-crystallized ligand conformation using different methods.
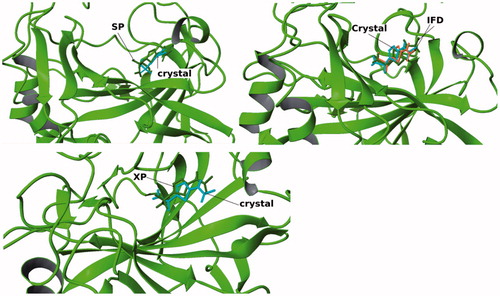
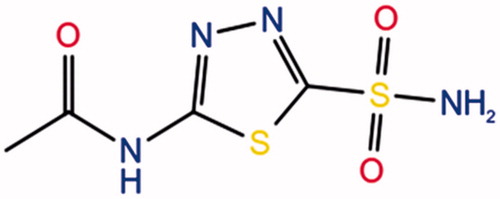
Table 1. 2D chemical structures and their corresponding docking scores for selected top-docking scored compounds.
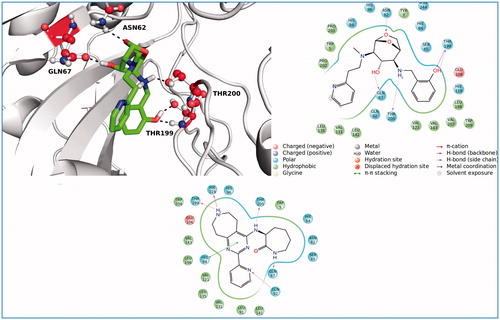
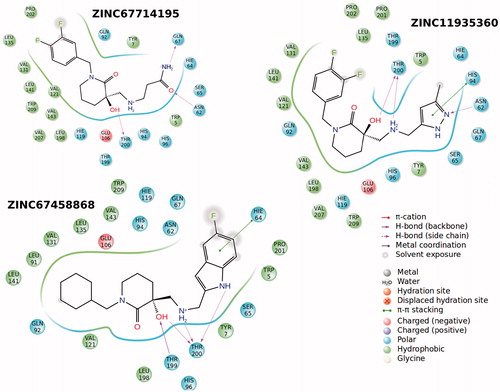
Figure 5. 3D and 2D ligand interaction diagrams for the ligands, ZINC20464003 (upper panel) and ZINC72421916 (lower panel).