Figures & data
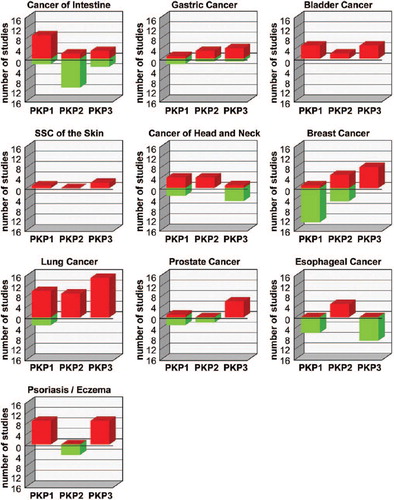
Table 1. PKP expression in cancer.