Figures & data
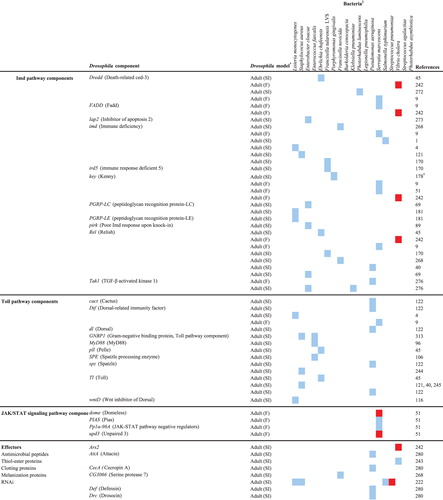
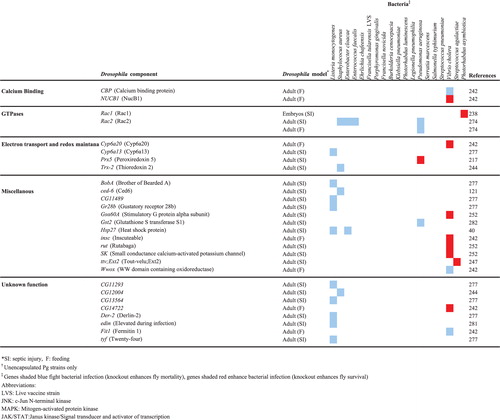
Table 1. Human bacterial pathogen genes that are involved in the infection of Drosophila