Figures & data
Table 1. Profile of the seven notifiable flaviviral species.
Figure 1. Overlap extension-polymerase chain reaction design algorithm.
This formulation was prepared to algorithmically plan and accurately calculate the desired product for OE-PCR. The process was initially done using manual algorithm whereby executable OE-PCR oligomers can be designed on the limitation of user-determined oligomer length. The platform made use of Linux (Ubuntu).
OE-PCR: Overlap extension-polymerase chain reaction.
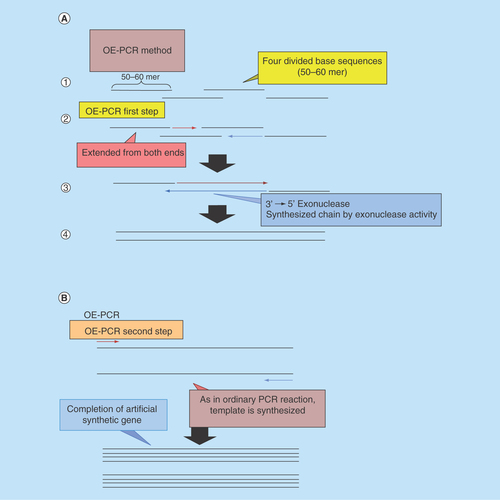
Figure 2. Diagram of the two-step cycle overlap extension-polymerase chain reaction (A: first step; B: second step).
1st PCR: For OE-PCR, 50–60mer oligomers were designed to have 19–23mer overlapping regions and make partial dimers (a). At the start of extension step of 1st PCR, DNA was synthesized by polymerase which has a 3′→5′ exonuclease activity (b). From the exonuclease activity, some oligomers were digested and replaced by synthesized DNA strand (c). Whole length of DNA was synthesized (d).
OE-PCR: Overlap extension-polymerase chain reactions.
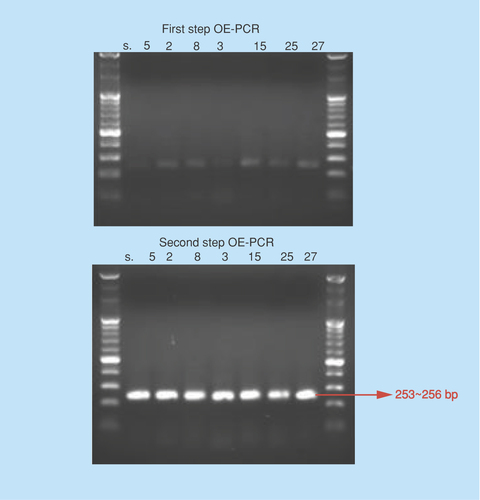
Figure 3. First and second step overlap extension-polymerase chain reaction showing prominent DNA bands.
Note the almost similar product sizes of 253–256 bp. Each of the four Dengue virus has 253 bp; Zika, yellow fever and Japanese encephalitis has 256 bp DNAs.
OE-PCR: Overlap extension-polymerase chain reactions.
Table 2. Genomic profile of newly synthesized dengue virus 1–4, Japanese encephalitis, yellow fever and Zika flaviviruses.
Table 3. Comparative features of oligomer design and assembly methods for overlap extension-polymerase chain reaction.
