Figures & data
Table 1. Physiological parameters used in the allometry scaling.
Table 2. Listing of species, doses, route, labeling, collection period and percent recovery in bile of various compounds/drugs that were subjected to interspecies scaling for allometry prediction.
Using the simple allometry (A, D & G), bile flow-rate corrected (B, E & H) and UDPGT corrected (C, F & I) methods of linagliptine, muraglitazar and odanacatib, respectively.
UDPGT: Uridine diphosphate glucuronosyltransferase.
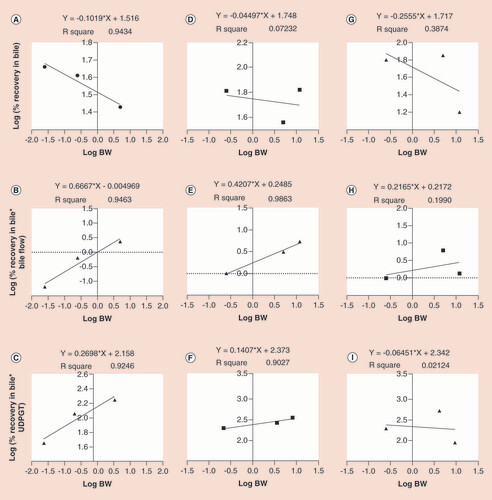
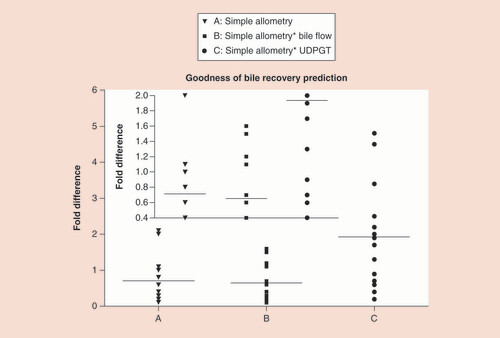
Table 3. Fold difference in the predicted and observed percent recovery of drug/drug-related amount of various compounds/drugs using different allometry methods.
Using (A) simple allometry, (B) bile flow-rate corrected allometry and (C) UDPGT activity corrected allometry (inset: an expanded scale is provided for easy visualization of the fold differences for the three allometry methods).
UDPGT: Uridine diphosphate glucuronosyltransferase.