Figures & data
Figure 1. C/EBPα is required for UVB-induced upregulation of p21 protein levels. (A) Immunoblot analysis of untreated control and C/EBPα siRNA treated Balb/MK2 keratinocytes and K5Cretg/+ and K5Cretg/+;C/EBPαflox/flox epidermal lysates. (B) Immunoblot analysis of lysates from Balb/MK2 keratinocytes treated with C/EBPα siRNA or control siRNA and collected at the indicated times after exposure to 5 mJ/cm2 UVB. (C) IHC staining for p21 in formalin-fixed paraffin-embedded sections of mouse skin from K5Cretg/+ and K5Cretg/+;C/EBPαflox/flox mice 4 h after treatment with 50 mJ/cm2 UVB. (D) Immunoblot analysis of p21, and C/EBPα from lysates isolated from K5Cretg/+ and K5Cretg/+;C/EBPαflox/flox mouse epidermis collected at the indicated times following exposure to 100 mJ/cm2 UVB. (E and F) Relative Cdkn1a (p21) mRNA levels in (E) Balb/MK2 cells treated with C/EBPα siRNA or control siRNA and (F) K5Cre and K5Cre;C/EBPαflox/flox in mouse epidermis collected at the indicated times following UVB. Data are expressed as the mean normalized to Gapdh ±S .D. (N ≥ 3). There were no statistically significant differences between control and C/EBPα-deficient keratinocytes in culture or in vivo as measured and calculated by Student's T test. (G) Immunoblot analysis of lysates of NHEK cells treated with C/EBPα siRNA or control siRNA and collected 10 h post 10 mJ/cm2 UVB. (H) Immunoblot analysis of Balb/MK2 treated with C/EBPα siRNA or control siRNA and collected 6 or 12 h after 25 μM MNNG treatment. (I) Immunoblot analysis of Balb/MK2 treated with C/EBPα siRNA or control siRNA and collected 8 h after exposure to 5 mJ/cm2 UVB.
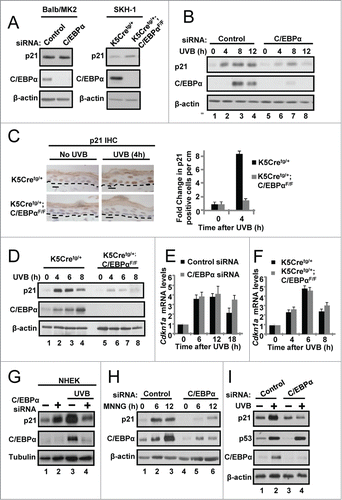
Figure 2. C/EBPα regulates nuclear p21 protein stability following UVB-induced DNA damage. (A) Immunoblot analysis of cytosolic and nuclear fractions prepared from siRNA treated Balb/MK2 keratinocytes collected 8 h post 5 mJ/cm2 UVB. (B) Immunoblot analysis of nuclear fractions prepared from siRNA treated Balb/MK2 collected at the indicated times post UVB. (C) Immunoblot analysis of p21 and C/EBPα in UVB and CHX treated nuclear extracts. Five h post 5 mJ/cm2 UVB cells were treated with 40 μg/mL CHX (t = 0) and collected at the indicated times. Values below the p21 image represent the fraction of p21 remaining at that time point as measure by densitometry normalized to β-actin. Values are also plotted to the right. (D) Immunoblot analysis of p21 and C/EBPα in CHX treated nuclear extracts. C/EBPα or control siRNA cells were treated with CHX. Values below the p21 image represent the fraction of p21 remaining at that time point as measure by densitometry normalized to β-actin. (E) Immunoblot analysis of p21 co-IP from nuclear extracts 6 h post 5 mJ/cm2 UVB.
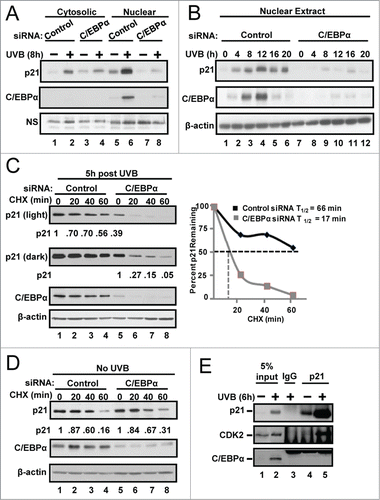
Figure 3. C/EBPα regulates CRL4Cdt2-mediated degradation of p21 in response to UVB. (A) Immunoblot analysis of control and C/EBPα siRNA nuclear extracts 8 h after 5 mJ/cm2 UVB. Ten μM MG132 was added 2 h before collection. (B) Immunoblot analysis of control and C/EBPα siRNA nuclear extracts 6 h after exposure to 5 mJ/cm2 UVB. Cells were treated with 800 nM MLN4924 30 min before UVB exposure. (C) Immunoblot analysis of control, C/EBPα, and Cdt2 siRNA nuclear extracts 6 h after 5 mJ/cm2 UVB. (D) Relative Cdt2 mRNA levels in control and C/EBPα siRNA 6 h following 5 mJ/cm2 UVB. Data are expressed as the mean normalized to Gapdh ±S .D. (N = 3). **= P value < 0.01 as calculated using Student's T test.
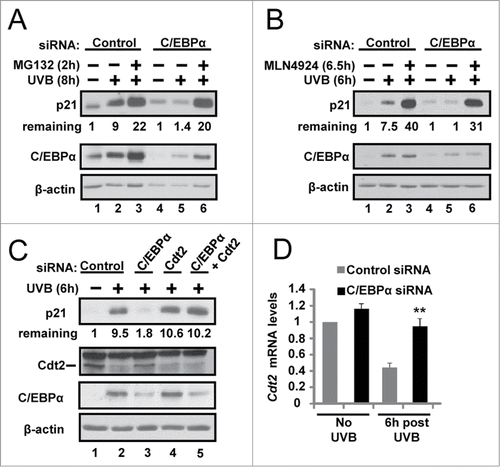
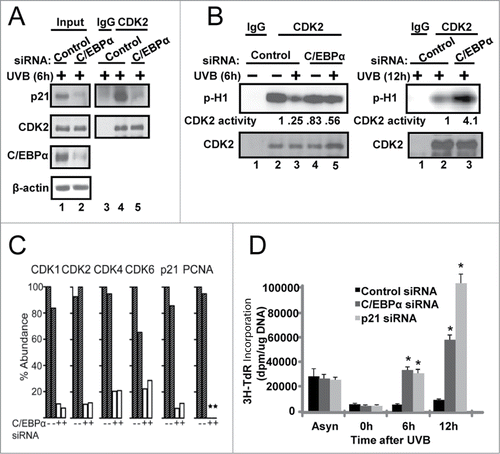
Figure 4. Failure to up-regulate nuclear p21 protein levels in UVB-exposed C/EBPα-deficient keratinocytes leads to functional consequences on cell cycle protein complexes, CDK2 activity and the G1/S checkpoint. (A) Immunoblot analysis of CDK2 co-IP from control and C/EBPα siRNA nuclear extracts 6 h following 5 mJ/cm2 UVB. (B) Phosphorylation of H1 by CDK2 immunoprecipitated from control and C/EBPα siRNA nuclear extracts 5 mJ/cm2 after UVB exposure. (C) Bar plot of peptide abundance for unique peptides for the proteins of interest across the 2 technical p21 IP replicates. Peptide abundances are expressed as a percent of the most abundant for a specific protein. **For PCNA, we could not confidently integrate a peptide signal in the siRNA replicates. A second peptide for each protein shows similar results (Fig. S3). (D)3H-thymidine incorporation in synchronized control and C/EBPα siRNA treated keratinocytes after 5 mJ/cm2 UVB exposure. Data represents the mean ± S.D., * = P value < 0.01 as calculated using Student's T test (N = 3) and are representative of 3 independent experiments.