Figures & data
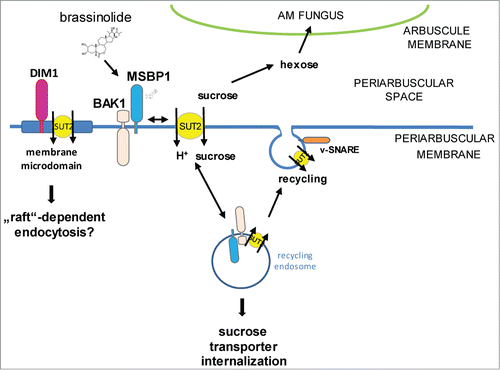
Figure 1. Mycorrhizal parameters of Oryza sativa Nipponbare wild type rice plants compared to brd2-1 mutant plants defective in BR biosynthesis 6 weeks after inoculation with Rhizophagus irregularis. F%: infection frequency, M%: absolute mycorrhizal colonization, m% relative mycorrhizal colonization, A% absolute arbuscule abundance, a% relative arbuscular abundance. Shown are mean values and standard deviations. Significant differences between WT and brd2-1 mutant plants are indicated by asterisks (Students-t-test, P ≤ 0.05; n = 4-6).
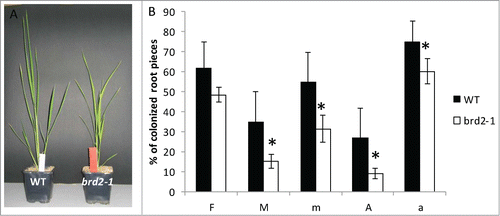
Figure 2. Quantitative real time PCR analysis of transcript levels of the DIM/DWARF1- (Os10g0397400) and OsSUT4-encoding (Q6YK44) genes relative to ubiquitin gene (XR_423446) transcript levels in Nipponbare wild type rice plants compared to the BR-synthesis defective brd2-1 mutant. Wild type plants were non-mycorrhizal (co) or inoculated with the AM fungi R. irregularis (myc). Shown are mean values and standard deviations. Significant differences between WT and brd2-1 mycorrhizal plants are indicated by asterisks (Students-t-test, P ≤ 0.05; n = 3-5).
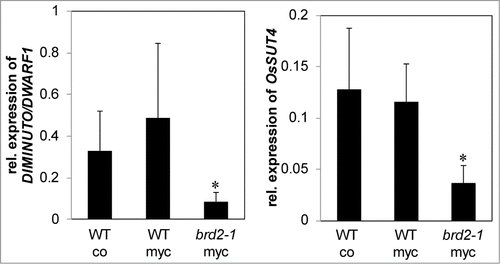
Figure 3. Hypothetical model describing the involvement of SlSUT2-interacting proteins in the subcellular localization of the sucrose transporter. SlSUT2 interacts with MSBP1, the LRR-receptor kinase BAK1-like and DIM1 that is associated to membrane microdomains. Endocytosis inhibits SlSUT2 transport activity, whereas recycling to the plasma membrane potentially via SNARE proteins is required for activation and sucrose retrieval from the periarbuscular space away from the AM fungus via SlSUT2. Brassinolide is assumed to affect mycorrhization via enabling MSBP1-SUT2 interaction for internalisation of SUT2 and inactivation its sucrose retrieval function.