Figures & data
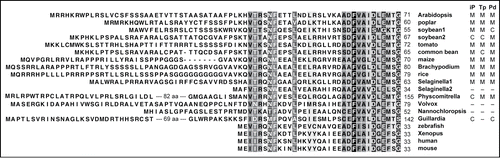
Figure 1. Comparison of the amino acid sequence of the N-terminal portion of poly(A) specific ribonucleases from various organisms. Highly conserved or relatively conserved amino acid residues are shaded in black and gray, respectively. The subcellular localization prediction results (M, mitochondria; C, plastids; –, other) from 3 prediction programs (iP, iPSORT; Tp, TargetP; Pd, Predotar) are summarized to the right of the species names. Accession numbers of PARN proteins: Arabidopsis, NP_175983.51; poplar, XP_002300380.2; soybean1, XP_003532241.1; soybean2, XP_003552994.1; tomato, XP_004244040.1; maize, NP_001169499.1; Brachypodium, XP_003581331.1; rice CAH66773.1; Selaginella1, XP_002969601.1; Selaginella2, XP_002987014.1; Physcomitrella, XP_001765586.1; Volvox, XP_002949475.1; Nannochloropsis, EWM30500.1; Guillardia, XP_005838739.1; zebrafish, AAQ97826.1; Xenopus, AAH73682.1; human, NP_002573.1; mouse, NP_083037.1.