Figures & data
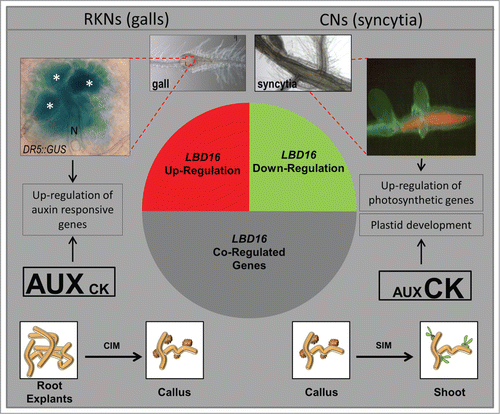
Figure 1. A simplified model showing the general results of in silico comparisons of the GCs and galls transcriptomes highlighting similarities to that of hormone-regulated genes and those co-expressed with LBD16. The most similar expression profile for the pool of genes co-regulated with LBD16 in galls and GCs was found in the transcriptome of Arabidopsis explants incubated on solid callus inducing media while those co-regulated in syncytia showed similar expression values to the transcriptomes of callus in the process of shoot regeneration indicated in the diagram. GUS activity of a DR5::GUS line on a 4 dpi gall showing intense staining in GCs. Syncytia show abundant chlorophyll fluorescence under fluorescence microscopy indicative of the presence of chloroplasts. N, nematode; all giant cells are marked with an asterisk. Drawings of callus in white boxes and their connections to the syncytia, GCs and galls microarray data are based on in silico analysis from the microarray experiments in Genevestigator released at August 2014 (Fig S1 and S2).