Figures & data
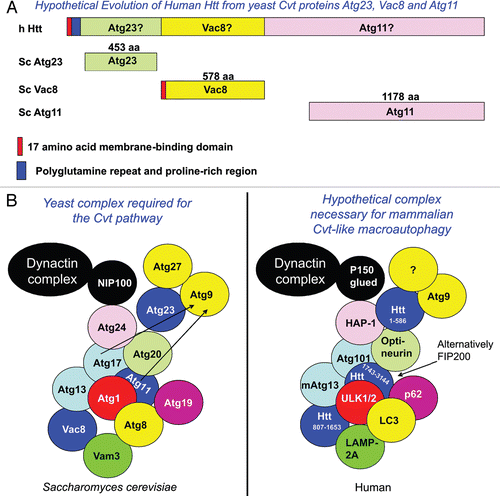
Table 1 Yeast Cvt pathway proteins and their hypothetical human functional homologs